Abstract
Free full text

Transcriptional super-enhancers connected to cell identity and disease
SUMMARY
Super-enhancers are large clusters of transcriptional enhancers that drive expression of genes that define cell identity. Improved understanding of the roles super-enhancers play in biology would be afforded by knowing the constellation of factors that constitute these domains and by identifying super-enhancers across the spectrum of human cell types. We describe here the population of transcription factors, cofactors, chromatin regulators and transcription apparatus occupying super-enhancers in embryonic stem cells and evidence that super-enhancers are highly transcribed. We produce a catalogue of super-enhancers in a broad range of human cell types, and find that super-enhancers associate with genes that control and define the biology of these cells. Interestingly, disease-associated variation is especially enriched in the super-enhancers of disease-relevant cell types. Furthermore, we find that cancer cells generate super-enhancers at oncogenes and other genes important in tumor pathogenesis. Thus, super-enhancers play key roles in human cell identity in health and disease.
INTRODUCTION
Transcription factors bind DNA regulatory elements called enhancers, which play key roles in the control of cell type-specific gene expression programs (Bulger and Groudine, 2011; Calo and Wysocka, 2013; Carey, 1998; Lelli et al., 2012; Levine and Tjian, 2003; Maston et al., 2006; Ong and Corces, 2011; Panne, 2008; Spitz and Furlong, 2012; Xie and Ren, 2013). A typical mammalian cell contains thousands of active enhancers, and it has been estimated that there may be ~1 million enhancers active in all human cells (Dunham et al., 2012; Heintzman et al., 2009; Thurman et al., 2012). It is important to further understand enhancers and their components because they control specific gene expression programs, and much disease-associated sequence variation occurs in these regulatory elements (Grossman et al., 2013; Lee and Young, 2013; Maurano et al., 2012).
The set of enhancers that control any one cell’s gene expression program is probably best defined in murine embryonic stem cells (ESCs). Co-occupancy of murine ESC genomic sites by the master transcription factors Oct4, Sox2 and Nanog is highly predictive of enhancer activity (Chen et al., 2008), and 8,794 enhancers have been identified in ESCs by using ChIP-Seq datasets for Oct4, Sox2 and Nanog (Whyte et al., 2013). A subset of these enhancers form 231 unusual enhancer domains at most genes that control the pluripotent state; these super-enhancers consist of clusters of enhancers that are densely occupied by five key ESC transcription factors and the Mediator coactivator (Whyte et al., 2013). There are many additional transcription factors, cofactors and chromatin regulators that contribute to the control of ESCs (Ng and Surani, 2011; Orkin and Hochedlinger, 2011; Young, 2011), and it would be instructive to know how these occupy enhancers and super-enhancers in ESCs. Similarly, it would be useful to know if super-enhancers are transcribed, because enhancer RNAs (eRNAs) have been proposed to contribute to enhancer activity (Lai et al., 2013; Lam et al., 2013; Li et al., 2013; Ling et al., 2004; Mousavi et al., 2013; Orom et al., 2010).
Super-enhancers are associated with key genes that control cell state in cells where they have been identified thus far, so identification of these domains in additional cell types could provide a valuable resource for further study of cellular control. We have generated a catalogue of super-enhancers in 86 human cell and tissue types. These super-enhancers are associated with genes encoding cell type-specific transcription factors, and thus identify candidate master transcription factors for many cell types that should prove useful for further understanding transcriptional control of cell state and for reprogramming studies. Using this catalogue, we find that DNA sequence variation associated with specific diseases is especially enriched in the super-enhancers of disease-relevant cells, suggesting that hypotheses regarding the role of specific cell types and genes in many diseases might be guided by knowledge of super-enhancers. Furthermore, tumor cells acquire super-enhancers at key oncogenes and at genes that function in the acquisition of hallmark capabilities in cancer, suggesting that these domains provide biomarkers for tumor-specific pathologies that may be valuable for diagnosis and therapeutic intervention. We discuss the implications of these observations for future study of disease.
RESULTS
Transcription factors in ESCs
Super-enhancers are clusters of enhancers, formed by binding of high levels of master transcription factors and Mediator coactivator, that drive high level expression of genes encoding key regulators of cell identity (Figure 1A) (Whyte et al., 2013). Five ESC transcription factors were previously shown to occupy super-enhancers (Oct4, Sox2, Nanog, Klf4, and Esrrb) (Whyte et al., 2013), but there are many additional transcription factors that contribute to the control of ESCs (Ng and Surani, 2011; Orkin and Hochedlinger, 2011; Young, 2011). We compiled ChIP-Seq data for 15 additional transcription factors in ESCs and investigated whether they occupy enhancers defined by Oct4, Sox2 and Nanog (OSN) co-occupancy (Whyte et al., 2013), (Table S1). The analysis showed that six additional transcription factors (Nr5a2, Prdm14, Tcfcp2l1, Smad3, Stat3 and Tcf3) occupy both typical enhancers and super-enhancers, and that all of these are enriched in super-enhancers (Figure 1B-E). Each of these factors has previously been shown to play important roles in ESC biology (Ng and Surani, 2011; Orkin and Hochedlinger, 2011; Young, 2011). In contrast, nine other transcription factors (c-Myc, CTCF, Zfx, Tbx3, YY1, Tfe3, Kap1/Zfp57, Smad1 and Ronin) were not similarly enriched in enhancers (Table S1), and instead occupied other regions of the genome such as promoter-proximal sites or sites that border topological domains (Figure S1A). It is particularly interesting that Smad3, Stat3 and Tcf3 are enriched in super-enhancer domains because these are transcription factor targets of the TGF-β, LIF and Wnt signaling pathways, respectively. Previous studies have shown that these transcription factors are recruited to enhancers formed by master transcription factors (Chen et al., 2008; Mullen et al., 2011), and evidence for enrichment of these factors at super-enhancers shows how these signaling pathways can converge on key genes that control ESC identity.
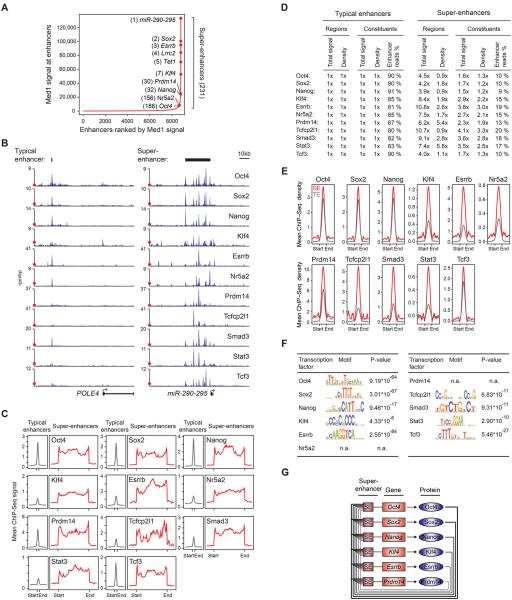
A) Distribution of Med1 ChIP-Seq signal at enhancers reveals two classes of enhancers in ESCs. Enhancer regions are plotted in an increasing order based on their input-normalized Med1 ChIP-Seq signal. Super-enhancers are defined as the population of enhancers above the inflection point of the curve. Example super-enhancers are highlighted along with their respective ranks and their associated genes.
B) ChIP-Seq binding profiles for the indicated transcription factors at the POLE4 and miR-290-295 loci in ESCs. Red dots indicate the median enrichment of all bound regions in the respective ChIP-Seq datasets, and are positioned at maximum 20% of the axis height. (rpm/bp: reads per million per base pair)
C) Metagene representations of the mean ChIP-Seq signal for the indicated transcription factors across typical enhancers and super-enhancer domains. Metagenes are centered on the enhancer region, and the length of the enhancer reflects the difference in median lengths (703bp for typical enhancers, 8667bp for super-enhancers). Additional 3kb surrounding each enhancer region is also shown.
D) Fold difference values of ChIP-Seq signal between typical enhancers and super-enhancers for the indicated transcription factors. Total signal indicates the mean ChIP-Seq signal (total reads) at typical enhancers and super-enhancers normalized to the mean value at typical enhancers. Density indicates the mean ChIP-Seq density at constituent enhancers (rpm/bp) of typical enhancers and super-enhancers normalized to the mean value at typical enhancers. Enhancer read % indicates the percentage of all reads mapped to enhancer regions that fall in the constituents of typical enhancer or super-enhancer regions.
E) Metagene representations of the mean ChIP-Seq density for the indicated transcription factors across the constituent enhancers within typical enhancers and super-enhancers. Each metagene is centered on enhancer constituents. Additional 2.5kb surrounding the constituent enhancer regions is also shown.
F) Table depicting transcription factor binding motifs enriched at constituent enhancers within super-enhancer regions, and associated p-values. The p-values of enrichment for the same motifs at typical enhancer constituents are listed in Supplemental Figure 1B.
G) Revised model of the core transcriptional regulatory circuitry of ESCs. The model contains an interconnected autoregulatory loop consisting of transcription factors that meet three criteria: 1) their genes are driven by super-enhancers, 2) they co-occupy their own super-enhancers as well as those of the other transcription factor genes in the circuit, and 3) they play essential roles in regulation of ESC state and iPSC reprogramming. The layout of the circuit model was adapted from (Whyte et al., 2013).
See also Figure S1.
To assess whether the 11 transcription factors that are enriched at super-enhancers contribute to super-enhancer formation by binding to known DNA sequence motifs, we analyzed the frequency of these binding motifs at super-enhancer regions. For all nine transcription factors for which binding motifs are available, we found that the cognate motif showed significant enrichment at super-enhancer constituents relative to background expectation (Figure 1F). These results suggest that the nine transcription factors contribute to super-enhancers by binding directly to their known DNA sequence motifs.
Previous studies have described a model of core transcriptional regulatory circuitry that includes Oct4, Sox2 and Nanog (Boyer et al., 2005). The evidence that these and additional ESC transcription factors form super-enhancers that drive genes essential for control of cell identity suggest a revised model of transcriptional regulatory circuitry for ESCs (Figure 1G). This model contains an interconnected autoregulatory loop like that originally proposed for Oct4, Sox2 and Nanog (Boyer et al., 2005), but consists of the additional ESC transcription factors that meet three criteria: 1) their genes are driven by super-enhancers, 2) they co-occupy their own super-enhancers as well as those of the other master genes and 3) they play important roles in regulation of ESC state and iPSC reprogramming.
RNA Polymerase II, co-factors and chromatin regulators
Super-enhancers are occupied by unusually high levels of the Mediator coactivator (Whyte et al., 2013). Previous studies have described the activities of RNA polymerase II and various cofactors, chromatin regulators and RNA at specific enhancers (Calo and Wysocka, 2013; Kagey et al., 2010; Lai et al., 2013; Lam et al., 2013; Li et al., 2013; Ling et al., 2004; Mousavi et al., 2013; Natoli and Andrau, 2012; Ong and Corces, 2011; Orom et al., 2010), so we used published and newly generated ChIP-Seq and RNA-Seq data to investigate how these components are associated with enhancers and super-enhancers across the ESC genome. The results indicate that RNA polymerase II, Mediator, cohesin, Nipbl, p300, CBP, Chd7, Brd4, and components of the esBAF (Brg1) and Lsd1-Nurd complexes are all enriched in super-enhancers relative to typical enhancers (Figure 2A-D; Table S1). RNA polymerase II can transcribe enhancers, producing non-coding RNAs that in some cases contribute to enhancer activity (Kim et al., 2010; Lam et al., 2013; Li et al., 2013; Natoli and Andrau, 2012; Sigova et al., 2013); we found that RNA polymerase II and RNA were highly enriched at super-enhancers relative to typical enhancers (Figure 2C).
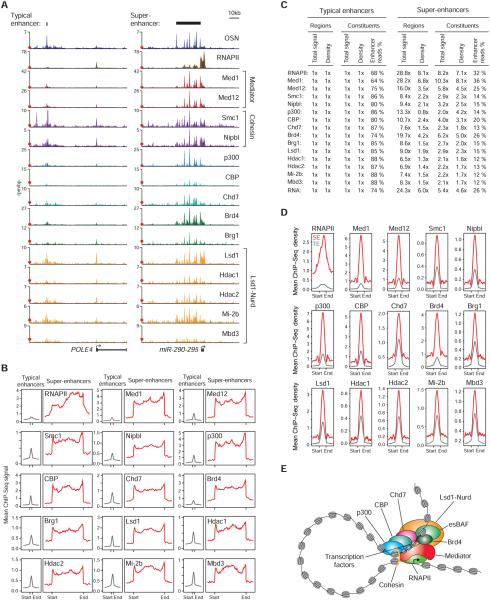
A) ChIP-Seq binding profiles for RNA Polymerase II (RNAPII) and the indicated transcriptional co-factors and chromatin regulators at the POLE4 and miR-290-295 loci in ESCs. OSN denotes the merged ChIP-Seq binding profiles of the transcription factors Oct4, Sox2 and Nanog, and serves as a reference. Red dots indicate the median enrichment of all bound regions in the respective ChIP-Seq datasets, and are positioned at maximum 20% of the axis height. (rpm/bp: reads per million per base pair)
B) Metagene representations of the mean ChIP-Seq signal for RNAPII and the indicated transcriptional co-factors and chromatin regulators across typical enhancers and super-enhancer domains. Metagenes are centered on the enhancer region, and the length of the enhancer reflects the difference in median lengths (703bp for typical enhancers, 8667bp for super-enhancers). Additional 3kb surrounding each enhancer region is also shown.
C) Fold difference values of ChIP-Seq signal between typical enhancers and super-enhancers for RNAPII and the indicated transcriptional co-factors and chromatin regulators, and RNA-Seq. Total signal indicates the mean ChIP-Seq signal (total reads) at typical enhancers and super-enhancers normalized to the mean value at typical enhancers. Density indicates the mean ChIP-Seq density at constituent enhancers (rpm/bp) of typical enhancers and super-enhancers normalized to the mean value at typical enhancers. Enhancer read % indicates the percentage of all reads mapped to enhancer regions that fall in the constituents of typical enhancer or super-enhancer regions. Reads mapped to exons were removed for the RNA-Seq analysis.
D) Metagene representations of the mean ChIP-Seq density for RNAPII and the indicated transcriptional co-factors and chromatin regulators across the constituent enhancers within typical enhancers and super-enhancers. Each metagene is centered on enhancer constituents. Additional 2.5kb surrounding the constituent enhancer regions is also shown.
E) Model showing RNAPII, transcriptional co-factors and chromatin regulators that are found in ESC super-enhancers. The indicated proteins are responsible for diverse enhancer-related functions, such as enhancer looping, gene activation, nucleosome remodeling and histone modification.
See also Figure S2.
It was notable that a broad spectrum of cofactors and chromatin regulators that are responsible for gene activation, enhancer looping, histone modification and nucleosome remodeling are especially enriched in ESC super-enhancers. The Mediator coactivator binds Nipbl, which loads cohesin, thus facilitating looping of enhancers to the promoters of their target genes (Kagey et al., 2010). The coactivator p300 is a histone acetyl-transferase, which is generally found at enhancer regions (Heintzman et al., 2007; Visel et al., 2009). CBP is a transcriptional co-activator that interacts with p300 and promotes synergy between enhancer components (Merika et al., 1998). Chd7 is a chromatin remodeler that also interacts with p300 and is often found at enhancers (Schnetz et al., 2010). Brd4, a member of the bromodomain protein family, binds to Mediator and acetylated histones, and is involved in regulation of transcriptional elongation by RNA polymerase II (Jang et al., 2005; Jiang et al., 1998). Brg1 is a subunit of the mammalian esBAF (SWI/SNF) complex, an ATP-dependent chromatin remodeler, which contributes to maintenance of pluripotency and self-renewal in ESCs (Ho et al., 2009a; Ho et al., 2009b). Lsd1, Hdac1, Hdac2, Mi-2b and Mbd3 are subunits of the Lsd1-Nurd complex, which possesses histone deacetylase, demethylase and nucleosome remodeling activities and contributes to enhancer decommissioning during differentiation (Denslow and Wade, 2007; Foster et al., 2010; Kaji et al., 2006; Kaji et al., 2007; Reynolds et al., 2012a; Reynolds et al., 2012b; Shi et al., 2004; Whyte et al., 2012).
Super-enhancers are occupied by an unusually large portion of the enhancer-associated RNA polymerase II and its associated cofactors and chromatin regulators. As measured by ChIP-Seq reads, between 12% and 36% of RNA polymerase II and the cofactors associated with all 8,794 enhancers were found within the 231 super-enhancers (Figure 2C). The evidence that a large fraction of these enhancer cofactors are associated with super-enhancers helps explain why these large domains produce relatively high levels of RNA (Figure 2C) and drive high-level expression of their associated genes when compared to typical enhancers (Whyte et al., 2013). The presence of high levels of RNA at super-enhancers is especially interesting in light of recent studies suggesting that enhancer RNA contributes to gene activation (Lai et al., 2013; Lam et al., 2013; Li et al., 2013; Ling et al., 2004; Mousavi et al., 2013; Orom et al., 2010), and evidence that the MYOD1 super-enhancer is transcribed into eRNAs that contribute to the transcriptional activation of MYOD1 in muscle cells (Mousavi et al., 2013).
ESC differentiation causes preferential loss of expression of super-enhancer - associated genes, which may be a consequence of the unusual vulnerability of super-enhancers to perturbation of their components (Loven et al., 2013; Whyte et al., 2013), (Figure S2). We speculate that this dual feature of super-enhancers – their ability to drive high level expression of key regulators of cell identity and their vulnerability to perturbation of their components – may facilitate cell state transitions during development.
Super-enhancers in many cell types
Because super-enhancers drive expression of genes that control and define cell identity, it would be useful to identify these elements and their associated genes in all human cells. However, the master transcription factors that might form super-enhancers are not known for most cell types and genome-wide binding data is limited for those that are known. We therefore explored the ability of various surrogate marks of enhancers (histone H3K27ac, H3K4me1, DNAse hypersensitivity, p300) to identify super-enhancers in ESCs (Creyghton et al., 2010; Heintzman et al., 2007; Neph et al., 2012; Rada-Iglesias et al., 2011; Shen et al., 2012; Visel et al., 2009). Of the marks available for a broad range of human samples, the histone H3K27ac modification was superior to the others in that it identified a large fraction of OSN-Mediator super-enhancers while minimizing excess sites (Figure S3A).
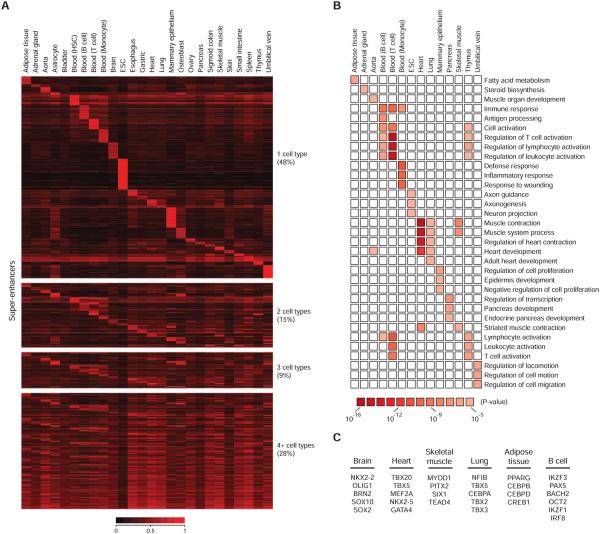
A) Heatmap showing the classification of super-enhancer domains across 26 human cell and tissue types. Color scale reflects the density of H3K27ac signal at the super-enhancer regions.
B) Gene Ontology terms for super-enhancer-associated genes in 13 human cell and tissue types with corresponding p-values.
C) Candidate master transcription factors identified in 6 cell types. All of these transcription factors were previously demonstrated to play key roles in the biology of the respective cell type or facilitate reprogramming to the respective cell type.
See also Figure S3.
We used H3K27ac ChIP-Seq data to create a catalogue of super-enhancers for 86 human cell and tissue samples (Figure 3; Table S2). A substantial portion of these super-enhancers are cell type-specific (Figure 3A, Figure S3B,C). Some of these domains overlap previously described locus control regions (LCRs), transcription initiation platforms (TIPs) and DNA methylation valleys (DMV), (Figure S3D), (Bonifer, 2000; Forrester et al., 1990; Grosveld et al., 1987; Koch et al., 2011; Tuan et al., 1985; Xie and Ren, 2013). Characterization of super-enhancer –associated genes by Gene Ontology analysis revealed that they are linked to biological processes that largely define the identities of the respective cell and tissue types (Figure 3B).
To gain further understanding of the transcriptional regulatory circuitry of cells, and to facilitate efforts to reprogram cells for regenerative medicine, it would be valuable to identify the master transcription factors that control all cell states (Cherry and Daley, 2012; Graf and Enver, 2009; Lee and Young, 2013; Zhou et al., 2008). Super-enhancers were previously identified in 5 murine cell types (ESC, myotubes, pro-B cells, Th cells and macrophages), and the genes encoding known master transcription factors in these cells were found to have associated super-enhancers (Whyte et al., 2013). We reasoned that candidate master transcription factors could be identified in most cells by identifying genes associated with super-enhancers that encode transcription factors and carried out this analysis in all the cells in this study. For those cells where master transcription factors have already been identified, this exercise captured the vast majority of these factors (Figure 3C). A catalogue of candidate master transcription factors for other cell types can be found in Supplemental Table 3. Prior studies of key transcriptional regulators suggest these candidates should be useful for deducing the transcriptional regulatory circuitry of many different human cells and for reprogramming studies.
Disease-associated DNA sequence variation in super-enhancers
Several recent studies suggest that much of disease-associated DNA sequence variation occurs in transcriptional regulatory regions defined by DNase hypersensitivity (Maurano et al., 2012; Vernot et al., 2012). We investigated the extent to which disease-associated DNA sequence variation occurs in enhancers and super-enhancers defined by histone H3K27ac. We compiled a list of 5,303 single nucleotide polymorphisms (SNP) linked to diverse phenotypic traits and diseases in 1,675 genome-wide association studies (GWAS), and investigated their distribution within enhancers and super-enhancers identified in the 86 human cell and tissue samples (Figure 4A, Table S4). We found that the majority of trait-associated SNPs occur in non-coding regions and that 64% of these occur within enhancer regions defined by H3K27ac (Figure 4A). Thus 64% of trait-associated non-coding SNPs occur in the ~33% of the genome covered by all enhancer regions defined by H3K27ac (Permutation test, P-value<10−4). The trait-associated SNPs were more enriched in super-enhancers than in typical enhancers (χ2 test, P-value<10−15) (Figure S4A), and for certain diseases, the enrichment in super-enhancers was particularly striking (Figure S4B). These results confirm that much of disease-associated DNA sequence variation occurs in transcriptional regulatory regions of the genome, indicate that most of this variation occurs in enhancers, and reveal that variation disproportionately impacts super-enhancer domains.
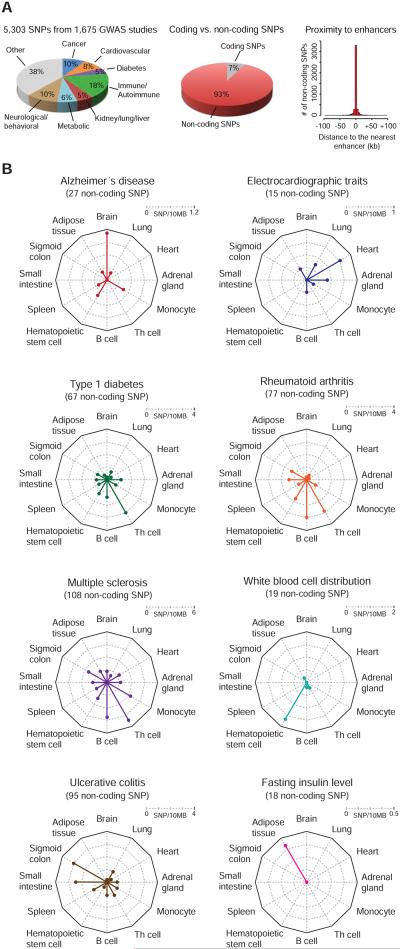
A) Catalogue of single nucleotide polymorphisms (SNP) linked to phenotypic traits and diseases in genome wide association studies (GWAS). (left) Pie chart showing percentage of SNPs associated with the highlighted classes of traits and diseases. (middle) Distribution of trait-associated SNPs in coding and non-coding regions of the genome. (right) Location of all non-coding trait-associated SNPs relative to all enhancers identified in 86 human cell and tissue samples. X-axis reflects binned distances of each SNP to the center of the nearest enhancer. SNPs located within enhancers are assigned to the 0 bin.
B) Radar plots showing the density of trait-associated non-coding SNPs linked to the highlighted traits and diseases, in the super-enhancer domains identified in 12 human cell and tissue types. The center of the plot is 0, and a colored dot on the respective axis indicates the SNP density (SNP/10MB sequence) in the super-enhancer domains of each cell and tissue type. Lines connecting the density values to the origin of the plot are added to improve visualization.
See also Figure S4.
If disease-associated SNPs occur disproportionately in super-enhancer domains, we would expect that SNPs associated with specific diseases would tend to occur in the super-enhancers of disease-relevant cells and not in those of disease-irrelevant cells. Indeed, for a broad spectrum of diseases, we found that disease-associated SNPs tend to occur in the super-enhancers of disease-relevant cells (Figure 4B; Table S4). This relationship was more pronounced for super-enhancers than typical enhancers (Figure S5). Since super-enhancers drive the expression of genes that control and define cell identity, these results suggest that altered expression of cell identity genes may often contribute to these diseases.
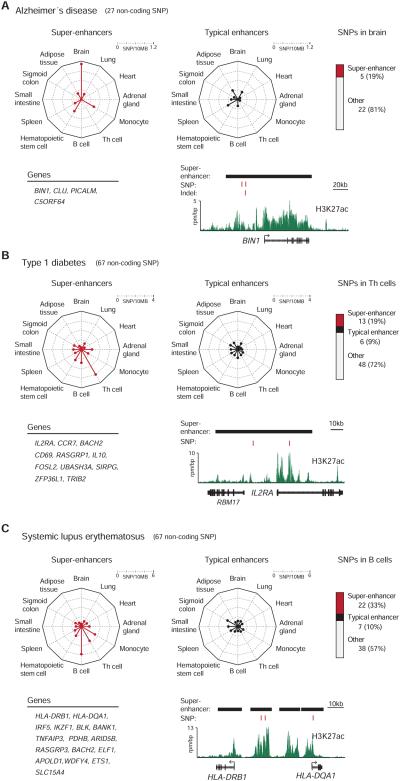
A) (top left) Radar plots showing the density of non-coding SNPs linked to Alzheimer’s disease (AD) in the super-enhancer domains and typical enhancers identified in 12 human cell and tissue types. The center of the plot is 0, and a colored dot on the respective axis indicates the SNP density (SNP/10MB sequence) in the super-enhancer domains or typical enhancers of each cell and tissue type. Lines connecting the density values to the origin of the plot are added to improve visualization. (top right) Distribution of non-coding SNPs linked to AD in the typical enhancers and super-enhancers of brain tissue. (bottom left) List of genes associated with AD SNP-containing super-enhancers in brain tissue. (bottom right) ChIP-Seq binding profile for H3K27ac at the BIN1 locus in brain tissue. The positions of AD-SNPs are highlighted as red bars, and the super-enhancers are highlighted as black bars above the binding profile. Indel rs59335482 (a three base pair insertion) is also highlighted. (rpm/bp: reads per million per base pair)
B) (top left) Radar plots showing the density of non-coding SNPs linked to type 1 diabetes (T1D) in the super-enhancer domains and typical enhancers identified in 12 human cell and tissue types. The center of the plot is 0, and a colored dot on the respective axis indicates the SNP density (SNP/10MB sequence) in the super-enhancer domains or typical enhancers of each cell and tissue type. Lines connecting the density values to the origin of the plot are added to improve visualization. (top right) Distribution of non-coding SNPs linked to T1D in the typical enhancers and super-enhancers of Th cells. (bottom left) List of genes associated with T1D SNP-containing super-enhancers in Th cells. (bottom right) ChIP-Seq binding profile for H3K27ac at the IL2RA locus in Th cells. The positions of T1D SNPs are highlighted as red bars, and the super-enhancers are highlighted as black bars above the binding profile.
C) (top left) Radar plots showing the density of non-coding SNPs linked to systemic lupus erythematosus (SLE) in the super-enhancer domains and typical enhancers identified in 12 human cell and tissue types. The center of the plot is 0, and a colored dot on the respective axis indicates the SNP density (SNP/10MB sequence) in the super-enhancer domains or typical enhancers of each cell and tissue type. Lines connecting the density values to the origin of the plot are added to improve visualization. (top right) Distribution of non-coding SNPs linked to SLE in the typical enhancers and super-enhancers of B cells. (bottom left) List of genes associated with SLE SNP-containing super-enhancers in B cells. (bottom right) ChIP-Seq binding profile for H3K27ac at the HLA-DRB1 and HLA-DQA1 loci in B cells. The positions of SLE SNPs are highlighted as red bars, and the super-enhancers are highlighted as black bars above the binding profile.
See also Figure S5.
Examples of disease-associated SNPs in super-enhancers
We focused further study on several diseases where SNPs occur in super-enhancers of disease-relevant cell types in order to gain further insights into the relationship between these SNPs, specific super-enhancers and their associated genes. The diseases we selected for further study included Alzheimer’s disease, type 1 diabetes, and systemic lupus erythematosus (Figure 5).
Alzheimer’s disease is a common form of dementia characterized by progressive neurodegeneration and loss of cognitive functions of the brain, and much of the genetic variation implicated in Alzheimer’s disease is associated with amyloid precursor protein, transmembrane proteins and apolipoprotein E4 (Bertram and Tanzi, 2008; Tanzi, 2012). The SNP catalogue contains 27 SNPs linked to Alzheimer’s disease and 5 of these occur in the super-enhancers of brain tissue (Figure 5A). Thus, ~19% (5/27) of all the Alzheimer’s disease SNPs occur in the 1.4% of the genome encompassed by brain tissue super-enhancers (Permutation test, P-value<10−2). Two SNPs occur in the super-enhancer associated with the gene BIN1 (Figure 5A), whose expression has recently been shown to be associated with Alzheimer’s disease risk (Chapuis et al., 2013). In this study, additional variation in BIN1 was associated with Alzheimer’s disease (Chapuis et al., 2013), involving a small insertion that occurs in the BIN1 super-enhancer identified here.
Type 1 diabetes is caused by T-cell mediated autoimmune destruction of pancreatic beta cells, and much of the genetic variation implicated in type 1 diabetes is associated with major histocompatibility antigens, interleukin-2 signaling, T-cell receptor signaling and interferon signaling (Bluestone et al., 2010; Noble and Erlich, 2012). The SNP catalogue contains 76 SNPs linked to type 1 diabetes, and 67 of these occur in non-coding sequences. The non-coding SNPs were particularly enriched in the super-enhancers of primary Th cells, with 13 occurring in the super-enhancer regions of genes with prominent roles in Th cell biology (Figure 5B). Again, it was striking that 19% (13/67) of all the Type 1 diabetes SNPs in noncoding regions occur in the 1.3% of the genome encompassed by Th cell super-enhancers (Permutation test, P-value<10−4).
Systemic lupus erythematosus (SLE) is a systemic autoimmune disease characterized by the loss of tolerance for self-antigens and production of excess amounts of serum autoantibodies, and most genetic variation associated with SLE involves major histocompatibility antigens and lymphocyte signaling pathways (Costa-Reis and Sullivan, 2013; Deng and Tsao, 2010). The SNP catalogue contains 72 SNPs linked to SLE and 67 of these occur in non-coding regions. Among the cell types examined here, the non-coding SNPs occur most frequently in B cell super-enhancers, with 22 SNPs occurring in the super-enhancer regions of 16 genes that play key roles in B cell biology (Figure 5C). Thus, 33% (22/67) of the SLE SNPs in non-coding regions occur in the 1.5% of the genome encompassed by B cell super-enhancers (Permutation test, P-value<10−4).
Similar enrichment of disease-associated variation in super-enhancers was observed for many additional diseases including rheumatoid arthritis, multiple sclerosis, systemic scleroderma, primary biliary cirrhosis, Crohn’s disease, Graves disease, vitiligo and atrial fibrillation (Table S4). This suggests that hypotheses regarding the role of specific cell types and genes in many diseases might be guided by knowledge of super-enhancers.
Super-enhancers in cancer
Super-enhancers associate with key oncogenes in several cancer cells (Loven et al., 2013). To gain further insights into the relationship between super-enhancers and cancer cell states, we used H3K27ac ChIP-Seq data to identify super-enhancers in 18 human cancer cells and identified their associated genes (Table S2). The data revealed that a remarkable spectrum of known oncogene drivers have associated super-enhancers in this set of cancer cells (Figure 6A; Table S2). These results suggest that super-enhancers may be useful for identifying key oncogenes in specific cancers.
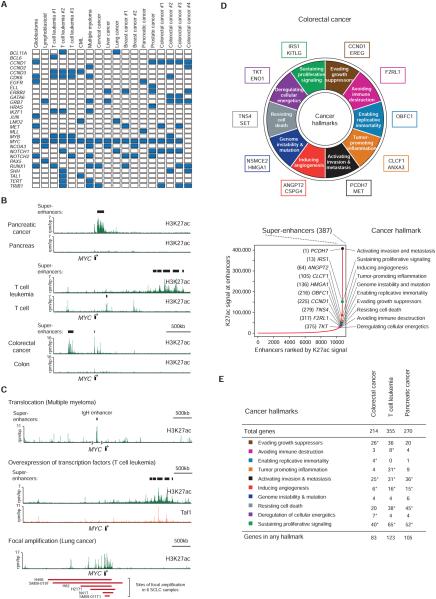
A) Selected genes associated with super-enhancers in the indicated cancers. Blue box indicates the gene being associated with a super-enhancer in the respective cancer. CML stands for chronic myelogenous leukemia.
B) Cancer cells acquire super-enhancers. ChIP-Seq binding profiles for H3K27ac are shown at the gene desert surrounding MYC in pancreatic cancer, T cell leukemia, colorectal cancer, and healthy counterparts. In colorectal cancer several regions in the 1MB window upstream of MYC were shown to interact with the MYC gene in colorectal cancer (Ahmadiyeh et al., 2010; Pomerantz et al., 2009). (rpm/bp: reads per million per base pair)
C) Chromosomal translocation, overexpression of transcription factors and focal amplification may contribute to super-enhancer formation in cancer. Displayed are ChIP-Seq binding profiles for H3K27ac and indicated transcription factors at the gene desert surrounding MYC in the indicated cancers. (top) A translocation event places the MYC gene proximal to an inserted IgH super-enhancer in multiple myeloma. (middle) Tal1 binding is observed at a distal super-enhancer in T cell leukemia. (bottom) Large H3K27ac domains are observed at the site of focal amplification in lung cancer. The red bars below the binding profiles indicate the genomic positions of focal amplification in six different samples, two of which (SM09-019T and SM09-11T1) are primary patient samples (Iwakawa et al., 2013).
D) Tumor-specific super-enhancers associate with hallmark cancer genes in colorectal cancer. (top) Diagram of the ten hallmarks of cancer adapted from Hanahan and Weinberg, 2011. Genes associated with super-enhancers in colorectal cancer but not in healthy colon samples were assigned to hallmark categories based on their functions and their previous implication in tumorigenesis. Prominent genes that associate with tumor-specific super-enhancers are highlighted at each cancer hallmark. (bottom) Distribution of H3K27ac signal across enhancers identified in colorectal cancer. Uneven distribution of signal allows the identification of 387 super-enhancers. Prominent genes associated with super-enhancers in colorectal cancer but not in healthy colon are highlighted with their respective super-enhancer ranks and cancer hallmarks they were assigned to.
E) Super-enhancers acquired by cancer cells associate with hallmark genes. Each cancer hallmark was assigned a Gene Ontology term, and the number of genes that are associated with acquired super-enhancers and are included in that GO term is displayed for each cancer. Asterisk denotes statistical significance above genomic expectation (hypergeometric test, p<0.05).
See also Figure S6.
Further analysis of super-enhancers in tumor cells and related healthy cells suggests that cancer cells acquire super-enhancers at oncogene drivers during the process of tumor pathogenesis (Figure 6B). For example, for multiple cancer cells, exceptionally large super-enhancers were found in the gene desert surrounding the c-MYC gene in the cancer cells but not in their healthy counterparts (Figure 6B). Furthermore, the super-enhancers formed in the MYC locus were tumor type specific (Figure 6B; Figure S6). These results are consistent with the model that cancer cells acquire cancer-specific super-enhancers at key oncogenes that are not present in their healthy counterparts.
DNA translocation, transcription factor overexpression and focal amplification occur frequently in cancer, and these mechanisms can account for the ability of cancer cells to acquire super-enhancers (Figure 6C). In Multiple myeloma, for example, tumor cells often have a translocation that places the 3′ IgH super-enhancer adjacent the MYC gene (Figure 6C). Overexpression of the TAL1 transcription factor in acute lymphoblastic leukemia (T-ALL) is associated with super-enhancer formation at another site in the MYC locus (Figure 6C). And focal amplification in lung cancer involves a large super-enhancer that spans the MYC gene and its normal regulatory elements (Figure 6C); tandem repeats of DNA segments can lead to the formation of clusters of enhancers.
Hanahan and Weinberg (2011) have proposed that cancer cells acquire a number of hallmark biological capabilities during the multistep process of tumor pathogenesis (Hanahan and Weinberg, 2011). We used these hallmarks as an organizing principle to investigate whether super-enhancer genes are associated with these biological capabilities in tumor cells. We identified super-enhancers that were acquired by cancer cells (not present in a healthy counterpart) and determined how their associated genes fit into the hallmarks. The results of such analysis with a colorectal cancer line revealed that over one-third of the super-enhancer genes have functions that are associated with a cancer hallmark (Figure 6D,E; Table S5). A similar analysis of two additional cancer lines confirmed that a large fraction of genes that acquire super-enhancers have hallmark functions (Figure 6E, Table S5). We conclude that cancer cells acquire cancer-specific super-enhancers at genes whose functions are associated with these hallmarks of cancer.
DISCUSSION
Super-enhancers were previously identified in a small number of cells, where they were shown to consist of large clusters of transcriptional enhancers formed by binding of master transcription factors and to be associated with genes that control and define cell identity (Loven et al., 2013; Whyte et al., 2013). We have extended our understanding of super-enhancers by identifying the population of transcription factors, cofactors, chromatin regulators and core transcription apparatus that occupy these domains in embryonic stem cells and by demonstrating that super-enhancers are highly transcribed. We have created a catalogue of super-enhancers for 86 different human cell and tissue types and shown that these are associated with genes encoding cell type-specific transcription factors and other components that play important roles in cell type-specific biology. Most importantly, we find that sequence variation associated with a broad spectrum of diseases is especially enriched in the super-enhancers of disease-relevant cell types and that cancer cells generally acquire super-enhancers at oncogenes and other genes that play important roles in cancer pathogenesis.
The enhancers and transcription factors that control embryonic stem cell state are probably better understood than those for any other cell type, making ESCs an excellent model for identifying components of super-enhancers (Ng and Surani, 2011; Orkin and Hochedlinger, 2011; Young, 2011). Several important insights were gained by studying how >35 transcription factors, cofactors, chromatin regulators and components of the core transcription apparatus occupy enhancers and super-enhancers in ESCs. All the enhancer-binding transcription factors are enriched at super-enhancers, with some so highly enriched that they distinguish super-enhancers from typical enhancers. The transcription factor targets of the TGF-β, LIF and Wnt signaling pathways are enriched in super-enhancers, suggesting how these signaling pathways converge on key genes that control ESC identify. Super-enhancers are occupied by a large portion of the enhancer-associated RNA polymerase II and its associated cofactors and chromatin regulators, which can explain how they contribute to high-level transcription of associated genes. Furthermore, the levels of RNA detected at super-enhancers vastly exceed those at typical enhancers, and recent evidence suggests that these eRNAs may contribute to gene activation (Lai et al., 2013; Lam et al., 2013; Li et al., 2013; Ling et al., 2004; Mousavi et al., 2013; Orom et al., 2010).
We have generated a catalogue of super-enhancers and their associated genes in a broad spectrum of human cell and tissue types. The super-enhancers tend to be cell type-specific and the genes associated with these elements tend to be cell type-specific in their expression and linked to biological processes that largely define the identities of the respective cell and tissue types. Genes that encode candidate master transcription factors and non-coding RNAs such as miRNAs are among those associated with super-enhancers. Thus, the super-enhancer catalogue should provide a valuable resource for further study of transcriptional control of cell identity and for reprogramming (Cherry and Daley, 2012; Graf and Enver, 2009; Lee and Young, 2013; Zhou et al., 2008).
Several recent studies suggest that much of disease-associated DNA sequence variation occurs in transcriptional regulatory regions defined by DNase hypersensitivity (Maurano et al., 2012; Vernot et al., 2012). We found that disease-associated SNPs occur in super-enhancers of disease-relevant cells and that this occurs more frequently for super-enhancers than typical enhancers. Since super-enhancers drive the expression of genes that control and define cell identity, these results suggest that altered expression of cell identity genes may often contribute to these diseases. These observations also suggest that hypotheses regarding the role of specific cell types and genes in many diseases might be guided in the future by knowledge of super-enhancers.
Cancer cells acquire super-enhancers at oncogene drivers during the process of tumor pathogenesis. Cancer cells appear to acquire super-enhancers through a variety of mechanisms, including chromosomal translocation of super-enhancers normally associated with other genes, focal amplification, or overexpression of an oncogenic transcription factor. The super-enhancers acquired by cancer cells are associated with a remarkably broad spectrum of oncogenes that have been described thus far in cancer. They are also associated with genes that function in the acquisition of hallmark capabilities in cancer (Hanahan and Weinberg, 2011). These results suggest that super-enhancers can provide biomarkers for cancer-specific pathologies that may be valuable for further understanding cancer biology, diagnosis and therapy.
EXPERIMENTAL PROCEDURES
Data analysis
ChIP-Seq datasets were aligned using Bowtie (version 0.12.9) to build version mm9 of the mouse genome or build version hg19 of the human genome. The GEO accession IDs for all analyzed datasets are listed in Supplemental Table 6.
Normalized read density of a ChIP-Seq dataset in any region was calculated as described (Whyte et al., 2013). ChIP-Seq reads aligning to the region were extended by 200bp and the density of reads per base pair (bp) was calculated. The density of reads in each region was normalized to the total number of million mapped reads producing read density in units of reads per million mapped reads per base pair (rpm/bp).
We used the MACS version 1.4.1 (Model based analysis of ChIP-Seq) (Zhang et al., 2008) peak finding algorithm to identify regions of ChIP-Seq enrichment over background. A p-value threshold of enrichment of 1e-9 was used for all datasets.
Enhancers were defined as regions of ChIP-Seq enrichment for transcription factors in murine ESCs, and H3K27ac in human cells. To accurately capture dense clusters of enhancers, we allowed enhancer regions within 12.5kb of one another to be stitched together.
The methods for identifying super-enhancers, and assignment of enhancers to genes are fully described in the Supplemental Information.
Supplementary Material
01
02
03
04
05
06
07
ACKNOWLEDGMENTS
We thank David Orlando and Charles Lin for bioinformatics support, Lee Lawton, Jamie Newman, Peter Rahl and Warren Whyte for sharing data, Nancy Hannett for assistance with data acquisition, Tom Volkert, Jennifer Love, Sumeet Gupta, and Jeong-Ah Kwon at the Whitehead Genome Technologies Core for Solexa sequencing, and members of the Young lab for helpful discussions. We acknowledge the Genome Centers at the Broad Institute and the University of California and San Diego for ChIP-Seq datasets produced in the framework of the ENCODE and NIH Roadmap Epigenome Projects, and the Rush Alzheimer’s Disease Center at Rush University Medical Center, Chicago (supported by P30AG10161, R01AG17917, R01AG36042, RC2AG36547). This work was supported by the National Institutes of Health grants HG002668 (RAY), CA109901 (RAY) and CA146445 (RAY, TL). R.A.Y. is a founder of Syros Pharmaceuticals.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- Ahmadiyeh N, Pomerantz MM, Grisanzio C, Herman P, Jia L, Almendro V, He HH, Brown M, Liu XS, Davis M, et al. 8q24 prostate, breast, and colon cancer risk loci show tissue-specific long-range interaction with MYC. Proceedings of the National Academy of Sciences of the United States of America. 2010;107:9742–9746. [Europe PMC free article] [Abstract] [Google Scholar]
- Bertram L, Tanzi RE. Thirty years of Alzheimer's disease genetics: the implications of systematic meta-analyses. Nature reviews Neuroscience. 2008;9:768–778. [Abstract] [Google Scholar]
- Bluestone JA, Herold K, Eisenbarth G. Genetics, pathogenesis and clinical interventions in type 1 diabetes. Nature. 2010;464:1293–1300. [Europe PMC free article] [Abstract] [Google Scholar]
- Bonifer C. Developmental regulation of eukaryotic gene loci: which cis-regulatory information is required? Trends in genetics : TIG. 2000;16:310–315. [Abstract] [Google Scholar]
- Boyer LA, Lee TI, Cole MF, Johnstone SE, Levine SS, Zucker JP, Guenther MG, Kumar RM, Murray HL, Jenner RG, et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell. 2005;122:947–956. [Europe PMC free article] [Abstract] [Google Scholar]
- Bulger M, Groudine M. Functional and mechanistic diversity of distal transcription enhancers. Cell. 2011;144:327–339. [Europe PMC free article] [Abstract] [Google Scholar]
- Calo E, Wysocka J. Modification of enhancer chromatin: what, how, and why? Molecular cell. 2013;49:825–837. [Europe PMC free article] [Abstract] [Google Scholar]
- Carey M. The enhanceosome and transcriptional synergy. Cell. 1998;92:5–8. [Abstract] [Google Scholar]
- Chapuis J, Hansmannel F, Gistelinck M, Mounier A, Van Cauwenberghe C, Kolen KV, Geller F, Sottejeau Y, Harold D, Dourlen P, et al. Increased expression of BIN1 mediates Alzheimer genetic risk by modulating tau pathology. Molecular psychiatry. 2013 [Europe PMC free article] [Abstract] [Google Scholar]
- Chen X, Xu H, Yuan P, Fang F, Huss M, Vega VB, Wong E, Orlov YL, Zhang W, Jiang J, et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell. 2008;133:1106–1117. [Abstract] [Google Scholar]
- Cherry AB, Daley GQ. Reprogramming cellular identity for regenerative medicine. Cell. 2012;148:1110–1122. [Europe PMC free article] [Abstract] [Google Scholar]
- Costa-Reis P, Sullivan KE. Genetics and epigenetics of systemic lupus erythematosus. Current rheumatology reports. 2013;15:369. [Abstract] [Google Scholar]
- Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA, et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proceedings of the National Academy of Sciences of the United States of America. 2010;107:21931–21936. [Europe PMC free article] [Abstract] [Google Scholar]
- Deng Y, Tsao BP. Genetic susceptibility to systemic lupus erythematosus in the genomic era. Nature reviews Rheumatology. 2010;6:683–692. [Europe PMC free article] [Abstract] [Google Scholar]
- Denslow SA, Wade PA. The human Mi-2/NuRD complex and gene regulation. Oncogene. 2007;26:5433–5438. [Abstract] [Google Scholar]
- Dunham I, Kundaje A, Aldred SF, Collins PJ, Davis CA, Doyle F, Epstein CB, Frietze S, Harrow J, Kaul R, et al. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74. [Europe PMC free article] [Abstract] [Google Scholar]
- Forrester WC, Epner E, Driscoll MC, Enver T, Brice M, Papayannopoulou T, Groudine M. A deletion of the human beta-globin locus activation region causes a major alteration in chromatin structure and replication across the entire beta-globin locus. Genes & development. 1990;4:1637–1649. [Abstract] [Google Scholar]
- Foster CT, Dovey OM, Lezina L, Luo JL, Gant TW, Barlev N, Bradley A, Cowley SM. Lysine-specific demethylase 1 regulates the embryonic transcriptome and CoREST stability. Molecular and cellular biology. 2010;30:4851–4863. [Europe PMC free article] [Abstract] [Google Scholar]
- Graf T, Enver T. Forcing cells to change lineages. Nature. 2009;462:587–594. [Abstract] [Google Scholar]
- Grossman SR, Andersen KG, Shlyakhter I, Tabrizi S, Winnicki S, Yen A, Park DJ, Griesemer D, Karlsson EK, Wong SH, et al. Identifying recent adaptations in large-scale genomic data. Cell. 2013;152:703–713. [Europe PMC free article] [Abstract] [Google Scholar]
- Grosveld F, van Assendelft GB, Greaves DR, Kollias G. Position-independent, high-level expression of the human beta-globin gene in transgenic mice. Cell. 1987;51:975–985. [Abstract] [Google Scholar]
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–674. [Abstract] [Google Scholar]
- Heintzman ND, Hon GC, Hawkins RD, Kheradpour P, Stark A, Harp LF, Ye Z, Lee LK, Stuart RK, Ching CW, et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature. 2009;459:108–112. [Europe PMC free article] [Abstract] [Google Scholar]
- Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA, et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature genetics. 2007;39:311–318. [Abstract] [Google Scholar]
- Ho L, Jothi R, Ronan JL, Cui K, Zhao K, Crabtree GR. An embryonic stem cell chromatin remodeling complex, esBAF, is an essential component of the core pluripotency transcriptional network. Proceedings of the National Academy of Sciences of the United States of America. 2009a;106:5187–5191. [Europe PMC free article] [Abstract] [Google Scholar]
- Ho L, Ronan JL, Wu J, Staahl BT, Chen L, Kuo A, Lessard J, Nesvizhskii AI, Ranish J, Crabtree GR. An embryonic stem cell chromatin remodeling complex, esBAF, is essential for embryonic stem cell self-renewal and pluripotency. Proceedings of the National Academy of Sciences of the United States of America. 2009b;106:5181–5186. [Europe PMC free article] [Abstract] [Google Scholar]
- Iwakawa R, Takenaka M, Kohno T, Shimada Y, Totoki Y, Shibata T, Tsuta K, Nishikawa R, Noguchi M, Sato-Otsubo A, et al. Genome-wide identification of genes with amplification and/or fusion in small cell lung cancer. Genes, chromosomes & cancer. 2013;52:802–816. [Europe PMC free article] [Abstract] [Google Scholar]
- Jang MK, Mochizuki K, Zhou M, Jeong HS, Brady JN, Ozato K. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Molecular cell. 2005;19:523–534. [Abstract] [Google Scholar]
- Jiang YW, Veschambre P, Erdjument-Bromage H, Tempst P, Conaway JW, Conaway RC, Kornberg RD. Mammalian mediator of transcriptional regulation and its possible role as an end-point of signal transduction pathways. Proceedings of the National Academy of Sciences of the United States of America. 1998;95:8538–8543. [Europe PMC free article] [Abstract] [Google Scholar]
- Kagey MH, Newman JJ, Bilodeau S, Zhan Y, Orlando DA, van Berkum NL, Ebmeier CC, Goossens J, Rahl PB, Levine SS, et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature. 2010;467:430–435. [Europe PMC free article] [Abstract] [Google Scholar]
- Kaji K, Caballero IM, MacLeod R, Nichols J, Wilson VA, Hendrich B. The NuRD component Mbd3 is required for pluripotency of embryonic stem cells. Nature cell biology. 2006;8:285–292. [Abstract] [Google Scholar]
- Kaji K, Nichols J, Hendrich B. Mbd3, a component of the NuRD co-repressor complex, is required for development of pluripotent cells. Development. 2007;134:1123–1132. [Abstract] [Google Scholar]
- Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, Harmin DA, Laptewicz M, Barbara-Haley K, Kuersten S, et al. Widespread transcription at neuronal activity-regulated enhancers. Nature. 2010;465:182–187. [Europe PMC free article] [Abstract] [Google Scholar]
- Koch F, Fenouil R, Gut M, Cauchy P, Albert TK, Zacarias-Cabeza J, Spicuglia S, de la Chapelle AL, Heidemann M, Hintermair C, et al. Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters. Nature structural & molecular biology. 2011;18:956–963. [Abstract] [Google Scholar]
- Lai F, Orom UA, Cesaroni M, Beringer M, Taatjes DJ, Blobel GA, Shiekhattar R. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature. 2013;494:497–501. [Europe PMC free article] [Abstract] [Google Scholar]
- Lam MT, Cho H, Lesch HP, Gosselin D, Heinz S, Tanaka-Oishi Y, Benner C, Kaikkonen MU, Kim AS, Kosaka M, et al. Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature. 2013;498:511–515. [Europe PMC free article] [Abstract] [Google Scholar]
- Lee TI, Young RA. Transcriptional regulation and its misregulation in disease. Cell. 2013;152:1237–1251. [Europe PMC free article] [Abstract] [Google Scholar]
- Lelli KM, Slattery M, Mann RS. Disentangling the many layers of eukaryotic transcriptional regulation. Annual review of genetics. 2012;46:43–68. [Europe PMC free article] [Abstract] [Google Scholar]
- Levine M, Tjian R. Transcription regulation and animal diversity. Nature. 2003;424:147–151. [Abstract] [Google Scholar]
- Li W, Notani D, Ma Q, Tanasa B, Nunez E, Chen AY, Merkurjev D, Zhang J, Ohgi K, Song X, et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature. 2013;498:516–520. [Europe PMC free article] [Abstract] [Google Scholar]
- Ling J, Ainol L, Zhang L, Yu X, Pi W, Tuan D. HS2 enhancer function is blocked by a transcriptional terminator inserted between the enhancer and the promoter. The Journal of biological chemistry. 2004;279:51704–51713. [Abstract] [Google Scholar]
- Loven J, Hoke HA, Lin CY, Lau A, Orlando DA, Vakoc CR, Bradner JE, Lee TI, Young RA. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell. 2013;153:320–334. [Europe PMC free article] [Abstract] [Google Scholar]
- Maston GA, Evans SK, Green MR. Transcriptional regulatory elements in the human genome. Annual review of genomics and human genetics. 2006;7:29–59. [Abstract] [Google Scholar]
- Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H, Reynolds AP, Sandstrom R, Qu H, Brody J, et al. Systematic localization of common disease-associated variation in regulatory DNA. Science. 2012;337:1190–1195. [Europe PMC free article] [Abstract] [Google Scholar]
- Merika M, Williams AJ, Chen G, Collins T, Thanos D. Recruitment of CBP/p300 by the IFN beta enhanceosome is required for synergistic activation of transcription. Molecular cell. 1998;1:277–287. [Abstract] [Google Scholar]
- Mousavi K, Zare H, Dell’orso S, Grontved L, Gutierrez-Cruz G, Derfoul A, Hager GL, Sartorelli V. eRNAs Promote Transcription by Establishing Chromatin Accessibility at Defined Genomic Loci. Molecular cell. 2013 [Europe PMC free article] [Abstract] [Google Scholar]
- Mullen AC, Orlando DA, Newman JJ, Loven J, Kumar RM, Bilodeau S, Reddy J, Guenther MG, DeKoter RP, Young RA. Master transcription factors determine cell-type-specific responses to TGF-beta signaling. Cell. 2011;147:565–576. [Europe PMC free article] [Abstract] [Google Scholar]
- Natoli G, Andrau JC. Noncoding transcription at enhancers: general principles and functional models. Annual review of genetics. 2012;46:1–19. [Abstract] [Google Scholar]
- Neph S, Vierstra J, Stergachis AB, Reynolds AP, Haugen E, Vernot B, Thurman RE, John S, Sandstrom R, Johnson AK, et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature. 2012;489:83–90. [Europe PMC free article] [Abstract] [Google Scholar]
- Ng HH, Surani MA. The transcriptional and signalling networks of pluripotency. Nature cell biology. 2011;13:490–496. [Abstract] [Google Scholar]
- Noble JA, Erlich HA. Genetics of type 1 diabetes. Cold Spring Harbor perspectives in medicine. 2012;2:a007732. [Europe PMC free article] [Abstract] [Google Scholar]
- Ong CT, Corces VG. Enhancer function: new insights into the regulation of tissue-specific gene expression. Nature reviews Genetics. 2011;12:283–293. [Europe PMC free article] [Abstract] [Google Scholar]
- Orkin SH, Hochedlinger K. Chromatin connections to pluripotency and cellular reprogramming. Cell. 2011;145:835–850. [Europe PMC free article] [Abstract] [Google Scholar]
- Orom UA, Derrien T, Beringer M, Gumireddy K, Gardini A, Bussotti G, Lai F, Zytnicki M, Notredame C, Huang Q, et al. Long noncoding RNAs with enhancer-like function in human cells. Cell. 2010;143:46–58. [Europe PMC free article] [Abstract] [Google Scholar]
- Panne D. The enhanceosome. Current opinion in structural biology. 2008;18:236–242. [Abstract] [Google Scholar]
- Pomerantz MM, Ahmadiyeh N, Jia L, Herman P, Verzi MP, Doddapaneni H, Beckwith CA, Chan JA, Hills A, Davis M, et al. The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nature genetics. 2009;41:882–884. [Europe PMC free article] [Abstract] [Google Scholar]
- Rada-Iglesias A, Bajpai R, Swigut T, Brugmann SA, Flynn RA, Wysocka J. A unique chromatin signature uncovers early developmental enhancers in humans. Nature. 2011;470:279–283. [Europe PMC free article] [Abstract] [Google Scholar]
- Reynolds N, Latos P, Hynes-Allen A, Loos R, Leaford D, O’Shaughnessy A, Mosaku O, Signolet J, Brennecke P, Kalkan T, et al. NuRD suppresses pluripotency gene expression to promote transcriptional heterogeneity and lineage commitment. Cell stem cell. 2012a;10:583–594. [Europe PMC free article] [Abstract] [Google Scholar]
- Reynolds N, Salmon-Divon M, Dvinge H, Hynes-Allen A, Balasooriya G, Leaford D, Behrens A, Bertone P, Hendrich B. NuRD-mediated deacetylation of H3K27 facilitates recruitment of Polycomb Repressive Complex 2 to direct gene repression. The EMBO journal. 2012b;31:593–605. [Europe PMC free article] [Abstract] [Google Scholar]
- Schnetz MP, Handoko L, Akhtar-Zaidi B, Bartels CF, Pereira CF, Fisher AG, Adams DJ, Flicek P, Crawford GE, Laframboise T, et al. CHD7 targets active gene enhancer elements to modulate ES cell-specific gene expression. PLoS genetics. 2010;6:e1001023. [Europe PMC free article] [Abstract] [Google Scholar]
- Shen Y, Yue F, McCleary DF, Ye Z, Edsall L, Kuan S, Wagner U, Dixon J, Lee L, Lobanenkov VV, et al. A map of the cis-regulatory sequences in the mouse genome. Nature. 2012;488:116–120. [Europe PMC free article] [Abstract] [Google Scholar]
- Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, Casero RA, Shi Y. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell. 2004;119:941–953. [Abstract] [Google Scholar]
- Sigova AA, Mullen AC, Molinie B, Gupta S, Orlando DA, Guenther MG, Almada AE, Lin C, Sharp PA, Giallourakis CC, et al. Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proceedings of the National Academy of Sciences of the United States of America. 2013;110:2876–2881. [Europe PMC free article] [Abstract] [Google Scholar]
- Spitz F, Furlong EE. Transcription factors: from enhancer binding to developmental control. Nature reviews Genetics. 2012;13:613–626. [Abstract] [Google Scholar]
- Tanzi RE. The genetics of Alzheimer disease. Cold Spring Harbor perspectives in medicine. 2012;2 [Europe PMC free article] [Abstract] [Google Scholar]
- Thurman RE, Rynes E, Humbert R, Vierstra J, Maurano MT, Haugen E, Sheffield NC, Stergachis AB, Wang H, Vernot B, et al. The accessible chromatin landscape of the human genome. Nature. 2012;489:75–82. [Europe PMC free article] [Abstract] [Google Scholar]
- Tuan D, Solomon W, Li Q, London IM. The “beta-like-globin” gene domain in human erythroid cells. Proceedings of the National Academy of Sciences of the United States of America. 1985;82:6384–6388. [Europe PMC free article] [Abstract] [Google Scholar]
- Vernot B, Stergachis AB, Maurano MT, Vierstra J, Neph S, Thurman RE, Stamatoyannopoulos JA, Akey JM. Personal and population genomics of human regulatory variation. Genome research. 2012;22:1689–1697. [Europe PMC free article] [Abstract] [Google Scholar]
- Visel A, Blow MJ, Li Z, Zhang T, Akiyama JA, Holt A, Plajzer-Frick I, Shoukry M, Wright C, Chen F, et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature. 2009;457:854–858. [Europe PMC free article] [Abstract] [Google Scholar]
- Whyte WA, Bilodeau S, Orlando DA, Hoke HA, Frampton GM, Foster CT, Cowley SM, Young RA. Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature. 2012;482:221–225. [Europe PMC free article] [Abstract] [Google Scholar]
- Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, Rahl PB, Lee TI, Young RA. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell. 2013;153:307–319. [Europe PMC free article] [Abstract] [Google Scholar]
- Xie W, Ren B. Developmental biology. Enhancing pluripotency and lineage specification. Science. 2013;341:245–247. [Abstract] [Google Scholar]
- Young RA. Control of the embryonic stem cell state. Cell. 2011;144:940–954. [Europe PMC free article] [Abstract] [Google Scholar]
- Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, et al. Model-based analysis of ChIP-Seq (MACS) Genome biology. 2008;9:R137. [Europe PMC free article] [Abstract] [Google Scholar]
- Zhou Q, Brown J, Kanarek A, Rajagopal J, Melton DA. In vivo reprogramming of adult pancreatic exocrine cells to beta-cells. Nature. 2008;455:627–632. [Europe PMC free article] [Abstract] [Google Scholar]
Full text links
Read article at publisher's site: https://doi.org/10.1016/j.cell.2013.09.053
Read article for free, from open access legal sources, via Unpaywall:
http://www.cell.com/article/S0092867413012270/pdf
Citations & impact
Impact metrics
Citations of article over time
Alternative metrics
Article citations
Epigenetics behind CD8<sup>+</sup> T cell activation and exhaustion.
Genes Immun, 14 Nov 2024
Cited by: 0 articles | PMID: 39543311
Review
Multiscale 3D genome rewiring during PTF1A-mediated somatic cell reprogramming into neural stem cells.
Commun Biol, 7(1):1505, 14 Nov 2024
Cited by: 0 articles | PMID: 39537822 | PMCID: PMC11561290
Discovery and prioritization of genetic determinants of kidney function in 297,355 individuals from Taiwan and Japan.
Nat Commun, 15(1):9317, 29 Oct 2024
Cited by: 0 articles | PMID: 39472450 | PMCID: PMC11522641
Large-scale analysis of the integration of enhancer-enhancer signals by promoters.
Elife, 12:RP91994, 28 Oct 2024
Cited by: 2 articles | PMID: 39466837 | PMCID: PMC11517252
The role of the dynamic epigenetic landscape in senescence: orchestrating SASP expression.
NPJ Aging, 10(1):48, 24 Oct 2024
Cited by: 0 articles | PMID: 39448585 | PMCID: PMC11502686
Review Free full text in Europe PMC
Go to all (2,120) article citations
Other citations
Data
Data behind the article
This data has been text mined from the article, or deposited into data resources.
BioStudies: supplemental material and supporting data
Similar Articles
To arrive at the top five similar articles we use a word-weighted algorithm to compare words from the Title and Abstract of each citation.
Master transcription factors and mediator establish super-enhancers at key cell identity genes.
Cell, 153(2):307-319, 01 Apr 2013
Cited by: 2393 articles | PMID: 23582322 | PMCID: PMC3653129
What are super-enhancers?
Nat Genet, 47(1):8-12, 01 Jan 2015
Cited by: 421 articles | PMID: 25547603
Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers.
Mol Cell, 58(2):362-370, 19 Mar 2015
Cited by: 312 articles | PMID: 25801169 | PMCID: PMC4402134
Enhancer RNAs: a missing regulatory layer in gene transcription.
Sci China Life Sci, 62(7):905-912, 26 Dec 2018
Cited by: 22 articles | PMID: 30593613
Review
Funding
Funders who supported this work.
NCI NIH HHS (4)
Grant ID: R01 CA146445
Grant ID: CA109901
Grant ID: CA146445
Grant ID: P01 CA109901
NHGRI NIH HHS (2)
Grant ID: HG002668
Grant ID: R01 HG002668
NIA NIH HHS (8)
Grant ID: R01 AG036042
Grant ID: R01 AG017917
Grant ID: RC2 AG036547
Grant ID: P30AG10161
Grant ID: R01AG17917
Grant ID: RC2AG36547
Grant ID: P30 AG010161
Grant ID: R01AG36042





