Abstract
Free full text

TGF-β/BMP signaling and other molecular events: regulation of osteoblastogenesis and bone formation
Abstract
Transforming growth factor-beta (TGF-β)/bone morphogenetic protein (BMP) plays a fundamental role in the regulation of bone organogenesis through the activation of receptor serine/threonine kinases. Perturbations of TGF-β/BMP activity are almost invariably linked to a wide variety of clinical outcomes, i.e., skeletal, extra skeletal anomalies, autoimmune, cancer, and cardiovascular diseases. Phosphorylation of TGF-β (I/II) or BMP receptors activates intracellular downstream Smads, the transducer of TGF-β/BMP signals. This signaling is modulated by various factors and pathways, including transcription factor Runx2. The signaling network in skeletal development and bone formation is overwhelmingly complex and highly time and space specific. Additive, positive, negative, or synergistic effects are observed when TGF-β/BMP interacts with the pathways of MAPK, Wnt, Hedgehog (Hh), Notch, Akt/mTOR, and miRNA to regulate the effects of BMP-induced signaling in bone dynamics. Accumulating evidence indicates that Runx2 is the key integrator, whereas Hh is a possible modulator, miRNAs are regulators, and β-catenin is a mediator/regulator within the extensive intracellular network. This review focuses on the activation of BMP signaling and interaction with other regulatory components and pathways highlighting the molecular mechanisms regarding TGF-β/BMP function and regulation that could allow understanding the complexity of bone tissue dynamics.
Introduction
Bone, a specialized form of connective tissue, is the main element of the skeletal system. Its formation is a very complex but finely orchestrated process in which bone morphogenetic protein (BMP) plays the major role in the regulation of osteoblast lineage-specific differentiation and later bone formation.1 More insight into the functions of BMP has been gained from in vivo experiments using transgenic animals to reflect the importance of BMP in osteogenesis. Studies on loss-of-function and gain-of-function mutations result in various bone-related abnormalities during development.2–4 BMP, discovered in 1965, is a unique extracellular multifunctional signaling cytokine belonging to the large transforming growth factor-beta (TGF-β) super family.5 The identification and isolation of BMP has created great attention for their potential role in bone regeneration at both heterotopic and orthotopic sites. Now, recombinant human-BMP2 and -BMP7 are commercially available and clinically have been used in various therapeutic interventions, such as bone defects, non-union fractures, spinal fusion, osteoporosis, and root canal surgery.6 About 60 TGF-β family members have been identified so far7 with two general branches: (i) BMP/growth and differentiation factor (GDF) and (ii) the TGF-β/activin/nodal branch/mullerian-inhibiting substance or anti-mullerian hormone.8 Besides being a regulator of bone induction, maintenance, and repair, BMP is also a critical determinant of the non-osteogenic embryological development of mammals. Disturbances in BMP regulation lead to the pathogenesis of a variety of diseases, such as osseous deformation (fibrodysplasia ossificans progressiva, FOP), autoimmune, cancer, and cardiovascular diseases.9
The evolutionary significance of the BMP family is highlighted by the conserved nature of the canonical TGF-β/BMP signaling over at least 700 million years that proves the vitality and importance of BMP for vertebrate physiology.10 The highly conserved canonical TGF-β/BMP linear signaling cascade engages the TGF-β/BMP ligands, two cell surface BMP receptors (BMPRs), and signal transducers, Smads.11 Mechanistically, phosphorylated C-terminus receptor-regulated-Smad (R-Smad), specific for the BMP pathway, interacts with various downstream proteins, including Runx2, which in turn can induce bone differentiation factors.12 On the other hand, non-canonical Smad-independent signaling (p38 mitogen-activated protein kinase, MAPK) pathway also implicates the Runx2 gene to control the mesenchymal precursor cell (MPC) differentiation.1 The coordinated activity of Runx2 and BMP-activated Smads is critical for bone formation. Furthermore, a large number of gene products and pathways pointed to promote osteoblastogenesis and bone formation for maintaining the stability of bone microenvironment.13 Bone dynamics maintain delicate interactions between TGF-β/BMP and other pathways, which are tightly regulated spatiotemporally, giving rise to the remarkable complexity, diversity, flexibility, and delicacy of TGF-β/BMP functions.14,15 Several findings explore different modes of cross-talk between BMP signaling and other major signaling pathways, namely Wnt, Hedgehog (Hh), Notch, and MAPK,16–18 in which Runx2 plays as a key integrator.19
Interconnected signaling is responsible for final target gene expressions required for osteogenesis. As perturbations in the signaling result in bone diseases, so there is a great potential for clinical applications of TGF-β/BMP molecules for the treatment of bone disorders. However, the outcomes of clinical applications of BMP largely rely on a precise design of cell therapies. So, the structure of BMP and its receptors is of immense interest due to their pathophysiological implications in osseous and non-osseous diseases. Hence, this review emphasizes on the structure of BMP and its receptors, and its role in integrated regulatory mechanisms in signaling. This review also highlights different modes of cross-talk between BMP signaling and the signaling pathways of MAPK, Wnt, Hh, Notch, and FGF, in which Runx2 is a key transcriptional regulator of osteoblast differentiation and bone formation. This knowledge of the integrated signaling pathways helps to enable the rational design of cell therapies to replace damaged or aged bone tissues.
BMP and its receptors
The human genome encodes more than 20 homodimeric or heterodimeric BMP ligands20 which can be divided into four distinct subfamilies, according to their amino acid sequence homology, structures, and functions: (i) BMP2 and 4; (ii) BMP5, 6, 7, 8a, and 8b; (iii) BMP9 and 10; and (iv) BMP12, 13, and 14.6 BMP, soluble local-acting signaling proteins can function in an endocrine, paracrine, and autocrine manner. BMP2, BMP4 through to BMP7, and BMP9 have been shown to induce intra-membranous and endochondral bone formation. Of these, BMP2, 6, and 9 appear to have important roles in the induction of mesenchymal cell differentiation into osteoblasts.21 However, not all members are truly osteogenic. For example, BMP7 is involved in kidney, eye, and limb development; BMP4, 7, and 14 are important for proper reproductive tissue development; BMP2, 3, and 7 contribute to cartilage regeneration; and BMP12 and 13 are required for normal tendon healing.22,23 Recent studies suggest that BMP6 and 9 may be more effective in promoting orthotopic bone formation when compared to BMP2 and 7.24,25Table 1 26–52 represents the biological functions of BMP, its receptors, effectors, and regulators.
Table 1
| Subtype (s) | Biological function (s) | Reference (s) | |
|---|---|---|---|
| Ligand(s) | BMP1 | Cleaves procollagens I, II, and III to produce fragments that self-associate into mature collagen fibrils in cartilage formation | 12 |
| BMP2/BMP2a | Induces bone morphogenesis and involved in heart formation | 26 | |
| BMP3A/Osteogenin | Negative regulator of bone morphogenesis, act as chemo-attractant, induces synthesis and secretion of TGF-β1 by monocytes | 27 | |
| BMP3B/GDF10 | Negative regulator of bone morphogenesis in embryonic stage but have a positive role in endochondral bone formation in mature animals | 28 | |
| BMP4/BMP2B | Involved in bone induction, cartilage, limb and kidney formation, tooth development and fracture repair | 29 | |
| BMP5 | Role in early developmental skeletal patterning, limb development and bone morphogenesis | 30 | |
| BMP6/Vrg1, Dvr6 | Active role in the induction of osteoblast lineage-specific differentiation of MPC | 31 | |
| BMP7/OP1 | Potential osteoinductive factor for epithelial osteogenesis. Role in bone homeostasis and calcium regulation. Also involved in kidney formation | 32 | |
| BMP8A/OP2 | Role in bone morphogenesis and in the maintenance of spermatogenesis | 27 | |
| BMP8B/OP3 | Found only in mice spermatogenesis to prevent of male adult germ cell apoptosis | 27 | |
| BMP9/GDF2 | Active role in the induction of osteogenesis from lineage specific differentiation of MPC and in mature osteoblasts. Additionally, function in hepatic reticulo-endothelial and nervous system | 33 | |
| BMP10 | Heart morphogenesis (trabeculation of embryonic heart) | ||
| BMP11/GDF11 | Role during embryogenesis in the development of dorsal root ganglia and dorsal lateral region of the spinal cord tissues. Also involved in axial skeleton patterning, eye development, pancreas development and kidney formation | 34 | |
| BMP12/CDMP3, GDF7 | Involved in tendon and ligament formation, and repair and development of sensory neurons | 27 | |
| BMP13/CDMP2, GDF6 | Involved in chondrogenesis and hypertrophy | 35 | |
| BMP14/CDMP1, GDF5 | Survival-promoting molecule for dopaminergic neurons. Enhances tendon healing and bone formation | 12 | |
| BMP15/GDF9b | Role in ovarian development and function | 36 | |
| BMP16/Nodal | Embryonic patterning | 12 | |
| BMP17/Lefty | Embryonic patterning | 12 | |
| BMP18/Lefty | Embryonic patterning | 12 | |
| Receptor(s) | ALK1/ACVRL1 | Mutations in ALK1 are associated with hemorrhagic telangiectasia type 2 (Rendu-Osler-Weber syndrome 2). And somatic mosaicism of ACVRL1 is linked to severe pulmonary arterial hypertension (PAH) | 37 |
| ALK2/ACVR1/ActRIA | Mutation in ACVR1/ALK2 is linked with FOP. Over-expression of mutant ALK2 lead FOP induced endothelial-to-mesenchymal transition and differentiation into chondrocytes, osteoblasts and adipocytes | 38 | |
| BMPRIA (ALK3) | Over-expression of ALK3/BMPR-IA in C2C12 cells induced osteoblastic phenotype in absence of exogenous BMP. Diseases associated with BMPR1A includes juvenile polyposis syndrome | 39 | |
| BMPRIB (ALK6) | Receptor for BMP7/OP1 and GDF5/BMP1. Missense mutations of the BMPR-IB gene in childhood lead to idiopathic PAH. BMPR-IB is also associated with chrondrodysplasia, acromesomelic, and brachydactyly type C | 40 | |
| BMPRII | BMPRII mutations are linked to PAH | 41 | |
| ActRIIA | Induces bone formation and improves skeletal integrity. Selectively required for Smad4 independent BMP7 evoked chemotaxis | 42 | |
| ActRIIB/ACVR2B | Plays a role in glucose and energy homeostasis | 43 | |
| TGF-βRI (ALK5) | Involves in perichondrial formation and function, cartilage integrity, and osteoblast differentiation during growth plate development | 44 | |
| TGF-βRII | TGF-β RII required for calcium homeostasis and bone metabolism | 45 | |
| ActRIB/ALK4 | ActRIB is required for chicken egg cylinder organization and gastrulation | 45 | |
| Effector(s) | Smad1 | Smad1 plays an essential role in bone development and postnatal bone formation. Precisely, ALK2 (R206H) with Smad1 induced osteoblastic differentiation | 46 |
| Smad2 | Transduces the signal of the TGF-β, and thus regulates multiple cellular processes, such as cell proliferation, apoptosis, and differentiation | 47 | |
| Smad3 | Regulates the genes important for cell fate, such as differentiation, growth and death. Smad3 and Smad4 form a complex to regulate TGF-β inducible transcription. Imbalance Smad3 activity has been implicated in the pathogenesis of scleroderma and cancer | 47 | |
| Smad4/DPC4 | Involves in many cell functions such as osteoblastosis, differentiation, apoptosis, gastrulation, and embryonic development. Mutation of Smad4 gene causes a juvenile polyposis syndrome, hereditary hemorrhagic telangiectasia (HHT), and myhre syndrome | 47 | |
| Smad5 | Increased Smad 5 and activated ALK2 cooperatively induce BMP signaling in FOP | 38 | |
| Smad6 | Role in axial and appendicular skeletal development | 48 | |
| Smad8/9 | Mutation found in patients with idiopathic PAH. | 41 | |
| Smad7 | Preventing the formation of Smad2/Smad4 complexes. A mutation located in Smad7 gene is a cause of susceptibility to colorectal cancer type 3 | 49 | |
| Integrator(s) | Runx1/CBFA2/PEBP2αB/AML1 | Requires for haematopoiesis. It also associated with several types of leukemia including M2 AML and breast cancer | 50 |
| Runx2/CBFA1/AML3and PEBP2αA | Runx2 is a key osteoblast-specific transcription factor that plays a central role in osteoblast differentiation, chondrocyte maturation, bone formation and remodeling. Expression is largely restricted to osteoblasts and mesenchymal condensations forming bones, cartilages and teeth | 51 | |
| Runx3/AML2/CBFA3/PEBP2αC | Required for neurogenesis, thymopoiesis and gastric epithelial cell proliferation | 52 |
BMP is synthesized by osteoblasts as 400–500 amino acid peptide precursors. The large precursor BMP consists a hydrophobic secretory leader (signal peptide at the N-terminus) and a pro-peptide sequence (non-conserved domain) joined to the mature region (C-terminus domain).6 The mature domain of each BMP has a region of seven conserved cysteine amino acids at their C-terminal which contains 100–140 amino acid residues, six of which are involved in forming a characteristic structural motif (a cysteine-knot with two finger-like double-stranded sheets).12 In general, the C-terminal active domain, by a subtilisin-like convertase cleavage at a consensus dibasic Arg-X-X-Arg site, is released to generate mature dimers by disulfide bonds and dimerization which is a prerequisite for bone induction.53 The N-terminal region controls the stability of the processed mature protein, and the downstream sequence adjacent to the cleavage site determines the efficiency of cleavage.54 BMP has sites for N and O glycosylation, which increases the stability and half-life of the protein, in addition to determining the specificity of receptor coupling.55 Nevertheless, only BMP2 and 4 possess a potential secondary cleavage site. BMP2 is a multifunctional growth factor and a homodimer of two 114 peptides subunits, and two receptor-binding motifs have been identified in the BMP2 ligand. The wrist epitope, comprising of residues from both BMP2 subunits, has a high-affinity binding site for BMPRIA, while the knuckle epitope consists of one subunit and weakly binds to BMPRII.45 On the other hand, the monomer (BMP7) is stabilized by three disulfide bonds, i.e., Cys-67-Cys-136 and Cys-71-Cys-138 forming a ring through which the third, Cys-38-Cys-104, passes.56 The cysteine knot constitutes the monomer core from which four strands of anti-parallel β-sheets emanate, forming two finger-like projections. Figure 1 summarizes the genomic and proteomic organization of BMP and its receptors.
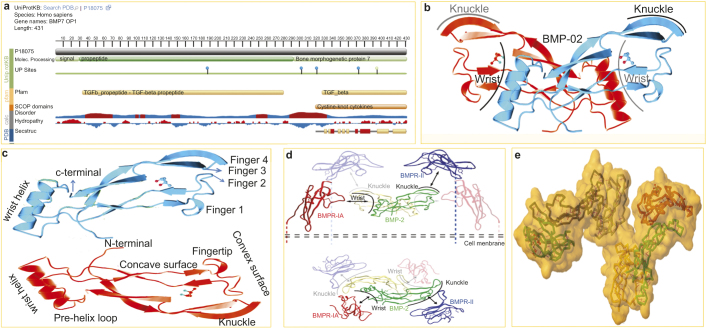
This picture represents the structure of BMP and its receptors. (a) The snapshot of BMP7/OP1 from uniprotKB/Swiss-Prot (Accession: P18075). Genomic location on 20q13 (chr20:55,746,015–55,841,178), size: 95,164 bp and code for 7 exon (UCSC Genome Browser ID: BMP7_HUMAN). OP1 is synthesized by osteoblasts as 431 amino acid peptides (Mw-49313 Da) precursor that is consist of a hydrophobic secretory leader (signal peptide at the N-terminus domain 1AA–29AA) and a pro-peptide sequences (non-conserved domain 30AA–292AA) joined to the mature region bone morphogenetic protein 7 (C-terminus domain 293AA–431AA). The displayed sequence is further processed into a mature form. Post-translational modifications (UP) sites are i.e.; glycosylation at Asn187, Asn302, Asn321 and Asn372, and Lys162 is the ubiquitination site, and 395AA site is for interchain disulphide bond formation. (b) Butterfly-shape crystal structure of human bone morphogenetic protein 2 (RCSB PDB entry 3BMP) from X-ray diffraction with resolution of 2.70 Å. Compound: 1 polymer, 1 ligand ((4S)-2-methyl-2, 4-pentanediol (C6H14O2)). The Jmol 3D cartoon shows two BMP2 subunit (Red and Blue) having wrist and knuckle epitopes for binding receptors. (c) A typical TGF-β/BMP consists of a cysteine knot motif with two pairs of antiparallel β-strands (fingers) extending from a α-helix (‘wrist’ region). The β-strands are curved to form both a concave and convex surface for receptor interaction. (d) Binding of BMP2 to BMPRII and BMPRIA receptors. Type I and type II receptors are glycoproteins of approximately 55 and 70 kDa, respectively. Side view (upper panel) and top view (lower panel) are shown. The ribbon diagram of hypothetical BMP2/BMPRIA/BMPRII ternary complex in the cell membrane is shown. BMP-2 has an elongated structure and binds to BMPRIA and BMPRII through wrist and knuckle epitopes, respectively. The structure of BMPRII was superposed onto that of ActR-II in the ternary complex containing BMP2, BMPRIA and ActR-II (Protein Data Bank entry 2GOO). Subunits of the BMP-2 dimer are shown in green and yellow. The extracellular domains of BMPR-IA and BMPR-II are shown in red and blue, respectively.45 (e) Structure of the ternary signaling complex of a TGF-β super family member. Ternary Complex of BMP2 binds to BMPR-IA and ActRII (PDB ID: 2GOO). The Jmol 3D shows the color subunit ternary BMP2 and BMPRs complex backbone in surface solvent accessible manner with yellow SS bonds and red H-bonds.
BMPRs are composed of three parts: a short extracellular domain, a single membrane-spanning domain, and an intracellular domain with the active serine/threonine kinase region.57 Three BMPRI (BMPRIA or ALK3, BMPRIB or ALK6, and ALK2: IA activin receptors) and three BMPRII (BMPRIIB, ACTRIIA, and ACTRIIB) are known to bind BMP. Out of the three BMPRI, ALK3 and ALK6 are structurally highly similar to each other, whereas ALK2 is less similar to the other two.51 One unique feature of BMPRI is the presence of a highly conserved TTSGSGSG motif (GS domain) in their cytoplasmic region that plays a key role in kinase activity.19 On the other hand, BMPRII has a unique, long C-terminal tail with 530 amino acids after the kinase domain.58 BMPRII has constitutively active kinases and BMPRI has inactive and inducible kinases. Recently, DRAGON, a glycosylphosphatidylinositol-anchored protein, was described as a co-receptor that directly interacts with ligands and receptors to facilitate BMP signaling.59 Again, Samad et al. also reported a second co-receptor: repulsive guidance molecule, a homologue of DRAGON, seems to be similarly involved in the BMP signaling pathway. The BMPRs were housed in a specific membrane domain of the caveolin-1 β (CAV1β) and clathrin-coated pits at cell surface.60
Normally, the BMP ligand initiates signaling by binding to and bringing together BMPRI and BMPRII receptors on the cell surface to form a ternary holocomplex.61 From the current understanding, heteromeric complexes of BMPRI and BMPRII are required for signal propagation. However, BMPRII is the primary binding site of the BMP ligand and phosphorylate of the BMPRII. And only then that BMPRIA signals are promoted to downstream substrates. It is believed that BMPRII does not actually bind the ligand but rather stabilizes and/or accelerates ligand binding to BMPRI. The specificity of the intracellular signals is mainly determined by BMPRI. Most typically, BMP2 and 4 bind to BMPRIA and BMPRIB, whereas BMP6 and 7 bind strongly to ALK2 and weakly to BMPRIB. BMPRII is specific for BMP, whereas ActRII and ActRIIB are shared by activins, myostatin, and BMP.62 The C-terminal tail of BMPRII has important functions in the regulation of cytoskeletal protein functions. BMPRII inhibits the ability of LMK1 to phosphorylate cofilin through interaction with its C-terminal tail and this inhibition is alleviated by BMP4.63Figure 1d,e illustrates the interaction of BMP and its receptor and receptor organization.
BMP signaling and implications in pathophysiology of bone formation
Both BMP and BMPRs are important not only for skeletal development and regeneration but also for the homeostasis of normal skeletal muscle mass. Recently it has been identified that the substitution of ″G″ with ″A″ in the activin A receptor type I/ACVR1 (ALK2) gene of juxta-membrane GS domain at position 617 (c.617 G>A) responsible for both familial and sporadic cases of FOP which is a rare autosomal dominant disorder that causes progressive heterotopic ossification (HO) in skeletal muscle tissue.64 Two clinical features define classic FOP: (i) malformation of the great toes and (ii) progressive HO in specific spatial patterns.65 The exact physiology behind FOP is still unclear, but the ectopic bone is thought to originate from MSCs which lie dormant in soft tissues and differentiate into osteogenic cells. Three states are involved in HO, namely a stimulating event, progenitor, or MSCs and an environment that allows osteogenesis.66 In humans, FOP causes metamorphosis of skeletal muscle and soft connective tissue into a second skeleton of heterotopic bone that mimic the patterns of normal embryonic skeletal formation, important differences are the lack of inflammation in embryonic skeletal induction and the relative absence of lymphocytic inflammatory cells in early fracture healing.67 A number of seminal discoveries provided evidence of profound dysregulation of the BMP signaling pathway in cells from patients who had FOP. The classic and invariable FOP phenotype of great toe malformations and progressive heterotopic endochondral ossification suggested that the primary molecular pathology involves the BMP signaling pathway.68 The recurrent mutation in ACVR1 causes an arginine-to-histidine substitution at p.R206H in ALK2 within the GS domain in FOP, making this one of the most highly specific disease-causing and first gain-of-function mutation in the human genome.69 We know that GS domain is the molecular switches of the type 1 receptor. Destabilization/autophosphorylation due to mutation of the GS domain promotes the correct positioning of glycine-Serine-rich loop and α-helix for kinase activation, consistent with an overactive BMP signaling pathway as the underlying cause of the ectopic chondrogenesis, osteogenesis, and joint fusion seen in FOP.68 The overexpression of BMP4 in lymphocytes has been reported in FOP patients, suggesting that BMP4 is one of the downstream target genes of BMP signaling in a positive feedback loop. Recently, additional genetic mutations have been found in ACVR1 in patients with HO and other skeletal abnormalities. The positions of the mutation in the GS domain are: p.L196P, p.P198-F199delinsL, p.R202I, and p.Q207E.70–72 Moreover, several mutations noticed in the serine/threonine kinase domain, such as p.R258S, p.G325A, p.G328E/R/W, p.G356D, and p.R375P.73–75Figure 2 shows that the FOP mutations map to the cytoplasmic GS and kinase domains of the ALK2 protein.
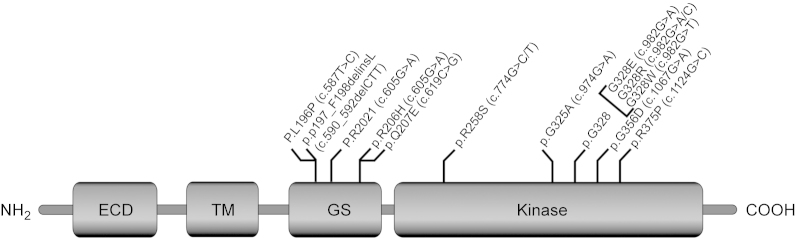
Schematic representation of the ALK2 protein showing the domain organization and gain of function FOP mutations map to the cytoplasmic GS and kinase domain.
Some of the mutations in the kinase domain have been suggested to be exposed at the interface with the GS domain and to be involved in the interaction with FKBP12/FKBP1A. The GS domain of all TGF-β/BMPRI is a critical site for binding and activation of pathway-specific Smad signaling proteins, and is a binding site of FKBP12, an inhibitory protein that prevents leaky activation of BMPRI in the absence of ligand.76 FKBP12 also recruits a Smad7-Smurf (Smad ubiquitylation regulatory factor)1 ubiquitin ligase that functions normally to regulate the abundance of the receptor at the membrane. Both leaky activation of BMP signaling and accumulation of BMP BMPRI at the cell membrane are seen in FOP cells, suggesting possible aberrant association with FKBP12 in FOP.77 The most likely possibility is that FKBP12 interactions with the GS domain become altered, leading to promiscuous ACVR1/ALK2 activity. However, exactly how the R206H mutation in ACVR1/ALK2 specifically perturbs BMP signaling in FOP remains a question but could involve dysregulation of BMPR functions or degradation/over activation of downstream signaling. This is of importance, as ACVR1 is a type 1 receptor for BMP, whose exogenous administration in soft tissue leads to ectopic bone formation.78 Thus, inhibition of mutant ACVR1 signaling in FOP patients represents an attractive therapeutic approach. Allele-specific RNA interference (ASP-RNAi), small chemical inhibitors of the kinase, including LDN-193189, Smad7 or dorsomorphin has been developed to prevent BMP signaling using a mutant ALK in FOP.79 However, there is still much to learn about the physiology and pathology at the interface of BMP and skeletal muscle, as FOP is, until recently, one of medicine's most elusive mysteries.
Smad (sma and mothers against decapentaplegic) protein: downstream effector of BMP signaling
The Smad is a group of intracellular proteins critical for transducing the signals from the cell surface to the nucleus upon binding of BMP ligands. There are three classes of Smad, namely (i) receptor regulated Smad (R-Smad), (ii) common mediator Smad (C-Smad), and (iii) antagonistic/inhibitory Smads (I-Smad).80 So far, Smad consist of three domains: an N-terminal mad-homology 1 (MH1) domain that carries nuclear localization signal and a DNA-binding region; a middle prolines, and serines/threonines-enriched linker domain that interacts with prolyl-isomerases and ubiquitin ligases; and a C-terminal MH2 domain that binds to BMPRI to mediate Smad signaling9,81. Generally, Smad forms a trimer which consists of two R-Smad and one C-Smad and acts as transcription factors that regulate the expression of certain osteoblastic genes.47 Nevertheless, the event of Smad/BMP interaction has been shown either an inhibitory or stimulatory function spatiotemporally through either Smad-dependent or Smad-independent pathways.12Figure 3 represents the proteomic organization of Smads and mechanism of interaction of Smad MH1 domain with DNA82.
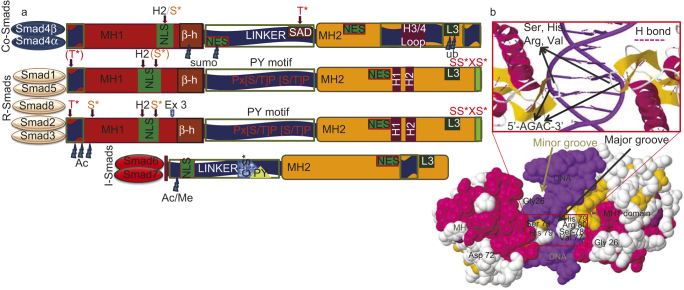
The proteomic organization of three subfamilies of Smads (C-Smad, R-Smad and I-Smad) and organization of a Smad MH1 domain with DNA. (a) All Smads information taken from PDB entry Smad>UniProt Gene list of Smads. The conserved N-terminal MH1 domain is in red, linker region in dark blue and the C-terminal MH2 domain in deep yellow. In the linker region the red PXS/TP (or S/TP) indicates the potential phosphorylation site for MAPKs ERK1/2, and the square indicates the PY (proline-tyrosine) motif that is recognized by the Hect/WW domain of Smurfs. The other domains and motifs are marked as follows: α-helix H1, H2, L3 and H3/4 loops, β–hairpin (β-h) that binds to DNA, the unique exon 3 of Smad2 (ex3), nuclear localization signal (NLS) and nuclear export signal (NES) motifs, Smad activation domain (SAD) at the linker-MH2 border. Sumoylation (sumo), ubiquitylation (ub), methylation (Me) and acetylation (Ac) sites are indicated with thundered heads. The unique SAD domain of Smad4 and the SSXS motif of R-Smads with asterisk indicating the phosphorylated serine residues. L3 loop and H1α helix in MH2 of Smad1, Smad5, Smad8 interacts with ALK1, ALK2, BMPR-IA, BMPR-IB by phosphorylation of Smad at C terminus. S*/T* indicate phosphorylatable serine and threonine residues. Brown S* phosphorylated by protein kinase C and by calmodulin-dependent kinase II. C-terminal red S*/S serines are phosphorylated by the type I receptor kinases. Different Smad interacting components, proteins and DNA with the specific functional domains of Smad are shown in the reviews by ten Dijke et al. (2000).82 (b) The 3D Java mol view of space filled secondary structure of a Smad MH1 domain bound to DNA (PDB entry 1MHD). Zoom in view represents the protein DNA interaction shows in red box. Highly conserved 11-residue β-hairpin loop recognizes major groove of DNA (dark blue) in a sequence-specific manner. In particular, β-hairpin loop in MH1 of Smad3, Smad4 interacts with 5″-AGAC-3″ termed Smad-binding elements.
Classical TGF-β/BMP signaling and its regulation by Smad
Among various Smad; R-Smad is the immediate downstream key molecule for the heterodimeric complex of two transmembrane serine/threonine kinase receptors to transduce BMP signal.83 Mechanistically, BMP ligand phosphorylate GS domain of BMPRIA and start the cascade for Smad. R-Smad has the structurally important L3 loop in the MH2 domain, which interacts with the type I receptor. R-Smad 1 and 5 become phosphorylated at the C-terminal serine residues of Ser-Ser-Val/Met-Ser sequence (SSXS motif).1 The specific phosphorylation of the R-Smad is controlled by two mechanisms of the receptor kinase complexes. First, a loop structure in the receptor kinase domain, the L4/5 loop, interacts with the L3 loop of an R-Smad protein. Second, R-Smad, such as Smad2 and Smad3, was phosphorylated by interacting with various cytoplasmic-anchored proteins beneath the cell membrane, like SARA (Smad Anchor for Receptor Activation) and cytoplasmic PML (promyelocytic leukemia) protein.84 Thus, upon BMP stimulation, BMPRII initiates the kinase activity of BMPRI leading to the phosphorylation of R-Smad in a complex cooperative process.
The phosphorylated R-Smad then disassociates from BMPRs, and forms a complex with C-Smad 4 which is imported to the nucleus. Then they interact with DNA-binding proteins or directly regulate transcriptional activity, either as monomers or in association with Smad4.6,85 In the nucleus they can regulate transcription of target genes by directly binding via their MH1 domain to specific DNA sequences depends on the formation of Smad complex with other DNA-binding proteins. The R-Smad/C-Smad heterodimer complex interacts with various transcription factors, co-activators and co-repressors to modulate gene expression.82,86 Smad complex binds the promotor regions of several BMP-responsive genes. Recently, the BMP responsive region in the Id1 promoter has been identified with two critical motifs, i.e., Smad-binding elements and GC-rich boxes (TGGCGCC).87,88 The sequence TGGCGCC (Bre7 motif) is located a short distance upstream of one or more Smad-binding elements motifs containing GTCTG. In the nucleus, Smads are also able to participate in histone modifications and chromatin remodeling.89 Runx2 in concert with R-Smad and/or Smad mediates specific signals to regulate osteoblasts formation and activate the expression of other osteogenesis specific target genes. The β-subunit of PEBP2 stabilizes Runx2 by preventing ubiquitin-dependent degradation.90 Upon activation of BMP signaling cascade, Runx2 and Smad physically interact to regulate the transcription of target genes co-operatively, and thereby induce osteoblast differentiation of MPCs. BMP does not directly induce the expression of Runx2 in mesenchymal cells, but it facilitates the expression of (Dlx5) in osteoblasts, and Dlx5 then induces expression of Runx2 in osteoprogenitor cells.91Figure 4 illustrates the classical TGF-β/BMP signaling and its regulation from BMP expression to Smad signaling including nuclear regulations.
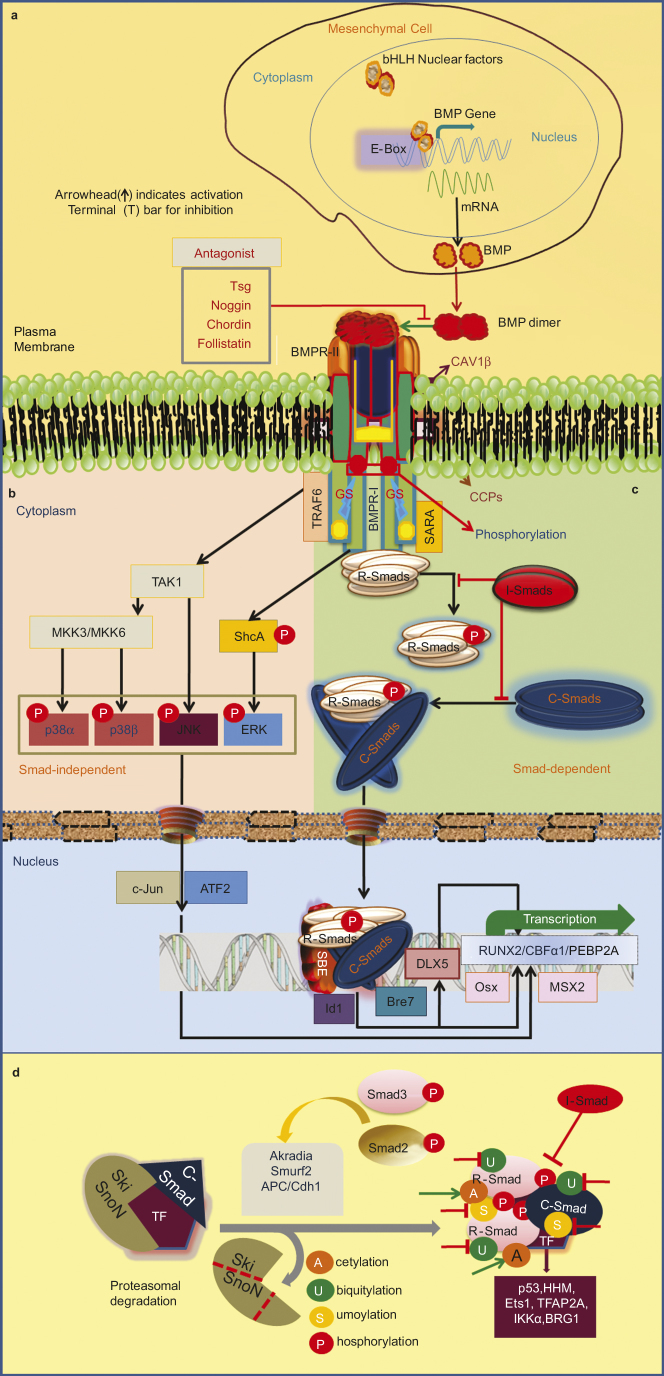
Schematic representation of BMP signaling and its regulation. (a) The basic helix-loop-helix (bHLH) proteins and its binding sequence (E-box) regulate the tissue-specific expression of the BMP gene. Osteogenic lineage-specific novel transcriptional factors can recognize the E-box. (b) Noncanonical Smad-independent signaling pathway. p38 activated MAPK pathway could converge at the Runx2 gene to control MPCs differentiation and TAK1 (TGF-β-activated kinase 1) signaling may also regulate bone formation. TGF-β-induced receptor heterotetramer recruit the ubiquitin ligase tumor necrosis factor α receptor associated factor 6 (TRAF6) to cytoplasmic domain. TRAF6 ubiquitylates and activates the catalytic activity of TAK1 and MAP3K7, leading to activation of the p38 and c-Jun N-terminal kinase (JNK) cascades, which regulate apoptosis and cell migration. The TGF-β type I receptor phosphorylates both serine and tyrosine residues in the SHCA (SHC1) adaptor, which then recruits the adaptor protein GRB2 and the Ras guanine exchange factor (GEF) son of seven less (SOS) in mammalian cells. This leads to activation of the Ras-Raf-MEK-Erk, MAPK JNK-c-Jun N terminal kinase, ATF-2 (activating transcription factor-2); p38 (p38 MAPK), SHCA (SH2 domain-containing sequence A), and Erk (extracellular signal regulated kinase). (c) Smad dependent pathway or the canonical BMP molecular signaling pathway. Type II BMP receptor and type I BMP receptor is housed in a specific membrane domain of CAV1β and clathrin-coated pits at cell surface. RGM acts as co-receptor of BMP signaling. Meanwhile BMP ligand dimer binds to BMPRII, BMPRI is cross-phosphorylated at GS site, and recruits R-Smad to the intracellular domain of the BMPR-I and initiates signal transduction via phosphorylation. Activated R-Smad then forms a heteromeric complex with C-Smad. This complex is translocated into the nucleus and interacts with several transcription factors such as Runx2/Cbfα1 (core binding factor alpha 1), Osx (Osterix), Dlx5, and Msx2 (msh homeobox homolog 2). These molecules mediate the transcription of related genes to induce osteogenesis. However, Smad complex binds the Id1 promoter that contains two critical motifs, i.e., SBEs and Bre7 motif. Noggin, twisted gastrulation (Tsg), and other antagonists bind to BMP ligands and block signaling. I-Smads reside in the nucleus, migrate to the cytoplasm and can negatively regulate BMP signaling by inhibiting signal transduction at several points. (d) Smad complex regulation inside the nucleus. The activated Smad complex interacts with a choice of Smad partners (transcription factors for instance BRG1 (Brahma-related gene 1), ETS1 (v-ets erythroblastosis virus E26 oncogene homolog 1), HHM (human homolog of Maid), IKKα (IκB kinase α), Smurf2, TFAP2A (transcription factor activating enhancer-binding protein 2α) and undergoes post-translational modifications. When the SNON/SKI are proteasomally degraded after being ubiquitylated by the ubiquitin ligases arkadia, Smurf2 or APC (anaphase-promoting complex)/CDH1 (ubiquitin ligase subunit) and thereby Smad target genes are inhibited. Nuclear R-Smads (e.g., Smad3) target the co-repressor SnoN for degradation via Smurfs or the APC that act as E3 ligases.9
Hh: modulator of BMP signaling
A number of findings have indicated that Hh proteins directly act on osteogenic precursor cells and osteoblasts to stimulate osteogenic differentiation.90 For example, Shh upregulates TGF-βII to inhibit hypertrophic chondrocyte differentiation during bone development. Whereas Shh/Gli2 induced BMP2 expression is responsible for osteoblast differentiation.91 Hence, during development, Shh and TGF-β/BMP pathways could directly regulate key components of each other. Both, Ihh and Shh have been shown to modulate BMP expression in cartilage and bone patterning throughout the axial, appendicular and facial skeleton.90 Shh is involved in fracture healing and bone maintenance. In the initial stages of fracture repair, the expression of Shh has been noticed in proliferating callus-forming cells in the periosteum.14 Additionally, the implantation of Shh-transduced cells increases the bone regeneration in a rabbit model of calvaria defects have also been found.92 In developing axial skeleton, sequential Shh and BMP signals are required for specification of a chondrogenic fate in presomitic tissue.93 Shh interact with the fibroblast growth factor 4 (FGF4) and BMP2 to induce the formation of osteoblast precursors and also involved in osteoblast differentiation by stimulating BMP2 promoter activity.19 The effects of Shh are mediated by transcription factor Gli2, a powerful activator of BMP2 gene expression, which is required for osteoblast differentiation.94 The expression of secreted phosphoprotein 1 (osteopontin) mediated via Gli2 and parathyroid hormone 1 receptor expressed via parathyroid hormone-like protein (PTHrP) signaling.91,95 PTHrP stimulates osteogenic cell proliferation. In developing long bones, Ihh stimulates the proliferation of chondrocytes but prevent differentiation at the growth plate in contract with BMP, and both protein acts on potential progenitor cells to promote osteoblast differentiation.40 Experimentally, Smad4 mutant mice showed in lessen of the expression of Ihh/PTHrP signaling in the growth plate.96 ALP activity also is regulated by Ihh and BMP synergistically. Moreover, upregulation of Ihh improve new bone formation at the osteogenic fronts through upregulation of BMP2.
Shh may increase mesenchymal proliferation via promotion of Msx2, and similarities are present between the Shh, Msx2, and BMP expression during neonatal craniofacial suture development.13 And Ihh induces PTCH and BMP2/4 expression, which results in intramembranous bone formation at the osteogenic fronts during intramembranous ossification.95 In sum, Ihh lies upstream of BMP2/4 signaling and may be a part of the regulatory network that controls BMP expression, thereby influencing bone formation. Discrete expression pattern of Shh versus Ihh in the developing skull indicates some possible differences in functions.97 Shh causes chondrocyte differentiation; Ihh induces differentiation of adjacent perichondrial cells into bone-forming osteoblasts.98 Our knowledge about the molecular nature of the BMP/Hh interaction is still limited; however, more sophisticated cross-talk between Hh and BMP is expected. Figure 5c,g summarizes hedgehog (Hh) signaling in osteoblasts proliferation, differentiation, as well as osteoclast formation.
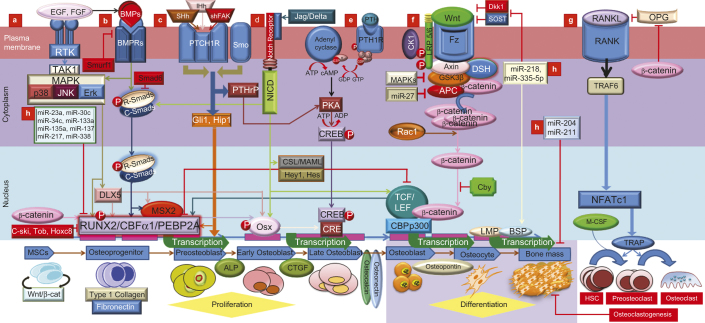
Schematic representation of lineage specific differentiation for MSCs in an exquisitely coordinated process with critical regulators and indicators are shown. This picture also highlights the different modes of cross-talk between TGF-β/BMP signaling and the major signaling pathways of MAPK, Wnt, Hh, Notch, and FGF in which Runx2 is a key transcriptional regulator of osteoblast differentiation and bone formation. (a) TAK1 signaling pathway regulates bone formation. Following BMP induction, MAPK pathways converge at the Runx2 and Dlx5 gene to control PMC differentiation. FGF stimulated TAK1 has been shown to increase ALP protein levels without changing Ocn and mRNA levels in the proliferation and differentiation of osteoblastic precursors. FGF may influence osteoblastogenesis at least partially through Runx2 modulation. FGF-2/EGF and BMP2 appear to be reciprocally regulated in osteoblasts, contributing to the balance of interacting signaling pathways in bone development and homeostasis. (b) BMP pathway, with their corresponding Smad proteins, and inhibitory proteins I-Smads (Smad6/7) and Smurf1 (Smad ubiquitination regulatory factor-1). Activated Smad regulates expression of transcriptional factors and transcriptional co-activators (Dlx5, Runx2 and Osx). Dlx5 the initial target of activated R-Smad which regulates Osx and Runx2/Cbfα1 to regulate osteoblast differentiation while Msx2 stimulates cell proliferation in a co-ordinatation with Shh. Ihh, Shh and BMP appear to be inter regulated in bone development. Osteoblasts specific markers, including the early osteogenic marker alkaline phosphatase (ALP), type I collagen, the late osteogenic markers osteocalcin (Oc) and osteopontin (Op), connective tissue growth factor (CTGF), inhibitor of DNA binding (Id) and CBFα1/Runx2. BMP2 exhibited increased mineralization by lime mineralization protein (LMP) and bone sialoprotein (BSP). Upon BMP9 stimulation of MSCs, CTGF was among the most up- regulated genes, especially during early stages of differentiation. Smurf1 targets type I BMP receptors and recognize bone-specific Runx2. Tob, Hox transcription factors inhibit BMP signaling as part of the negative feedback circuit. Ski onco-protein also can block BMP signaling. (c) Hedgehog-induced osteoblastogenesis occurs through Runx2. Hh binds cell surface receptor patched to relieve patched mediated suppression of Smoothened (Smo). Then Smo activate to stabilize the transcription factor Gli2, which induces transcription of Gli1 and other Hh target genes. Ihh signaling is required for early osteoblastogenesis, likely through modulation of Runx2, Osx, and ALP. And, Ihh and BMP2 appear to be reciprocally regulated in osteoblasts, contributing to the balance of interacting signaling pathways. (d) Notch signaling is important for MSC differentiation into osteoblasts. When Notch interacts with membrane bound ligands delta or jagged on the surface of neighboring cells, the Notch receptor liberating the notch intracellular domain (NICD) which binds to CSL. CSL then recruits the co-activator mastermind-like (MAML) for transcription of Hey/Hes to inhibit Runx2. In addition, NICD can interact directly with Runx2 protein to repress terminal osteoblastic differentiation. Signaling can also regulate either Smad dependent BMP signaling or Wnt-induced osteogenesis through the up-regulation of RANKL and OPG, indicating the cross-talk between osteoblasts and osteoclasts could be mediated by Notch signaling. (e) PTH binding activates PTH1R to stimulate several downstream effectors. Transcriptional factor cAMP response element binding protein (CREB) mediates PTH signaling in osteoblasts. (f) Canonical Wnt/β-catenin pathway increases bone mass through a number of mechanisms including renewal of stem cells, stimulation of preosteoblast replication, induction of osteoblastogenesis, and inhibition of osteoblast and osteocyte apoptosis. Upon intracellular accumulation of β-catenin, and enter into nucleus to interact with the transcription factor T-cell factor/lymphoid enhancer factor (TCF/LEF) for expressing of bone lineage genes such as Dlx5 and Osx. Wnt has also been linked to Runx2. Runx2 gene promoter contains a Wnt-responsive TCF regulatory element, and both β-catenin and TCF1 are recruited to the Runx2 locus. Wnt signaling and LRP5/6 coreceptor activity can be blocked by the sclerostin (SOST) and Dickkopf (Dkk), leading to a decrease in bone mass. Increasing the ratio of osteoprotegerin (OPG) to RANKL, β-catenin represses osteoclastogenesis. BMPRIA signaling upregulates Sost expression primarily through Smad-dependent signaling, while it upregulates DKK1 through Smad-dependent and non-Smad-dependent signaling. Chibby (Cby), the endogenous antagonist, interrupts the binding of β-catenin to transcriptional factors Tcf/Lef-1. (g) Sonic hedgehog (Shh) signaling is activated in osteoblasts and regulated their proliferation, differentiation, as well as osteoclast formation, via focal adhesion kinase (FAK) signaling. Shh expression is positively correlated with phosphorylated FAK Tyr to increase mesenchymal proliferation and suture mesenchyme thickness via promotion of Msx2, and similarities are present between the expression of Shh, Msx2, and BMP expression during neonatal craniofacial suture development. Shh signaling indirectly induced osteoclast differentiation by upregulating osteoblasts expression of PTHrP, which promoted receptor activator of nuclear factor kappa B ligand (RANKL) expression via PKA and its target transcription factor CREB. (h) Micro-RNAs that participate in stimulation or inhibition of osteoblast differentiation and their target genes are represented. Among these, miR-133 targets the transcription factor Runx2, a known target of BMP/Smad signaling that promotes osteoblast differentiation. Whereas, miR-218 and miR-335-5p can down regulates Dkk and Sost to play role in the period from osteocyte to bone mineralization. On the other hand, miR-135a, miR-23a, miR-133a, miR-137, miR-217 target Smad5 and Runx2 to inhibit BMP signaling.
Wnt/β-catenin: mediator of TGF-β/BMP signaling
The central function of Wnt/β-catenin pathway is the regulation of the stability of the β-catenin. In absence of Wnt ligand, β-catenin phosphorylated for proteasome mediated degradation that is controlled by the so-called ″β-catenin destruction complex″ which comprised of axin, adenomatous polyposis coli (APC), glycogen synthase kinase (GSK) 3 and casein kinase 1α).99 But in presence of Wnt, β-catenin accumulates and translocates to nucleus to initiate downstream gene expression. When Wnt ligand binds to its receptor frizzled (Fz) and co-receptor LRP5/6, the signal is transduced to the β-catenin destruction complex through an intracellular dishevelled protein. And thereby it leads to axin downregulation, GSK3 inactivation and β-catenin stabilization.100 More interestingly, inactivation of GSK3-β and stabilization of β-catenin reduces Smad1 ubiquitination.101 In the developing skeletal system, the Wnt/β-catenin signaling regulates the differentiation of progenitor's cells into osteoblast.18 Wnt/β-catenin pathway also help to increase the bone mass through a number of mechanisms including renewal of stem cells, stimulation of preosteoblast replication, induction of osteoblastogenesis, and inhibition of osteoblast and osteocyte apoptosis as well.102 Several Wnts (Wnt4, Wnt5a, Wnt9a, and Wnt16) are expressed in the perichondrium of developing long bones.103 However, the best-defined venue of BMP/Wnt cross-talk is in the nucleus, whereas the Smad/β-catenin/Lef (lymphocyte enhancer factor) complex regulates a group of shared target genes in a synergistic manner.
In Xenopus, BMP2/4 regulate Wnt-8 expression and cooperatively involved in the mesoderm patterning,104 whereas BMP2 downregulates Wnt7a and β-catenin in chicken embryonic mesenchymal cells in a p38-dependent manner, leading to enhanced chondrogenesis. In preosteoblastic cells, BMP2 modulates nuclear β-catenin signaling through stimulation of Wnt, LRP, and Fz receptors expression in mesenchymal cells, suggesting the interaction between Wnt and BMP2 signaling during osteoblastic differentiation.105 HesR1, TCF, LEF and Herp2 are the transcription factors specifically expressed in osteoblast cells by BMP treatment, and are also involved in Wnt signaling.106 Recent studies in differentiating osteoblasts indicate that Wnt pathway induction stabilizes β-catenin and increases TCF/LEF-dependent gene expression in parallel with β-catenin-independent complex formation between TCF4 and Runx2. Activation of either Runx2 or TCF4 co-enhances TCF and Runx2 activity increases TGF-βRI expression.107 Canonical Wnt/β-catenin pathway interacts with Runx2 as a critical mediator of BMP9-mediated osteogenic signaling.24 BMP9 stimulation of MSCs recruits β-catenin and Runx2 to the osteocalcin promoter.1 Wnt3a and BMP9 also synergized to induce ALP activity in MSCs, ectopic bone formation and matrix mineralization while BMP9-induced ALP activity inhibited by Frzb overexpression or β-catenin knockdown.24 The Wnt/β-catenin signaling pathway is activated during fracture repair. In early pluripotent mesenchymal stem cells, Wnt/β-catenin signaling needs to be precisely regulated to facilitate the differentiation of osteoblasts; by contrast, β-catenin is not needed for chondrocyte differentiation.108 Once mesenchymal stem cells are committed to the osteoblast lineage, activation of Wnt/β-catenin signaling enhances bone formation. Figure 5f highlights the cross-talk between BMP signaling and the signaling pathway of Wnt/β-catenin.
MicroRNAs act as a TGF-β/BMP signaling regulator
Activation of several signaling pathways, including TGF-β/BMP, Wnt/β-catenin, as well as transcription factors which are tightly regulated by differential expression of miRNAs.109 Osteoblast differentiation has been found tightly controlled by several regulators including miRNAs,12,110 and miRNAs can regulate expression of genes during differentiation of MSCs towards osteoblastic cells, resulting in bone formation. With regard to the regulation of bone formation, a growing number of miRNAs are found to be expressed in the developing skeletal system of metazoan. Besides the emerging role of miRNAs during embryo skeletogenesis, miRNA-dependent modulation of gene function can alter skeletal phenotypes across individuals and also within same individual over time.111
Recent evidence suggests that miRNAs might have either positive or negative regulatory function in osteoblast differentiation. For example, miR-218 facilitates differentiation of osteoblast into the final stage of forming mineralized tissue by inhibiting ERB1 (TOB1) and sclerostin (SOST).112 And miR-2861 promotes BMP2 induced ST2 osteoblast differentiation by repressing histone deacetylase 5 (HDAC5) expressions.113 BMP2 stimulates acetylation of Runx2 protein by p300 resulting inhibition of Smurf1-mediated degradation of Runx2 and increases the Runx2 transactivation activity. HDAC5 proteins can deacetylate Runx2 protein, and thus Runx2 protein is vulnerable to Smurf-mediated degradation. Homeobox A2 (Hoxa2), another repressor of Runx2 is targeted by miR-3960 leading to enhanced osteogenesis in BMP2 induced ST2 stromal cells.114 miR-27 promotes osteoblast differentiation by inhibiting APC gene expression, consequently activates Wnt/β-catenin signaling through accumulation non-phosphorylated β-catenin that translocates from cytoplasm to nucleus and then associates with TCF/LEF to induce target gene expression.19 Kapinas et al.115 and Zhang et al.116 have shown that miR-29a and miR-335-5p promotes osteoblast differentiation by downregulating the inhibitors of canonical Wnt/β-catenin signaling such as dickkopf-related protein 1 (Dkk1) and secreted Fz-related protein (sFRP2). MiR-208 enhances BMP2 induced pre-osteoblast differentiation by targeting the transactivation of osteopontin, Runx2, type I procollagen and PTHrP.117 Another miRNA, namely miR-138, regulates the osteogenic differentiation of human MSCs in vivo.118 Microenvironment determines the role of miRNAs for regulation of osteoblast differentiation. Liu et al.119 reported that miR-17 has a role for stimulation or inhibition of osteoblast differentiation that depends on the microenvironment. Table 2120–138 represents a list of microRNA with their regulatory functions in Osteoblast differentiation.
Table 2
| miRNA (s) | Function(s) | Reference (s) |
|---|---|---|
| miR-31 | Controls cytoskeleton organization in osteoclasts for optimal bone resorption activity | 120 |
| miRNA-34c | Regulates Notch signaling during bone development, and decrease osteoporosis | 121 |
| miR-210 | Acts as a positive regulator of osteoblastic differentiation by inhibiting the TGF-β/activin signaling through inhibition of AcvR1B | 122 |
| miR-20a | Increases BMP signaling by targeting antagonists of the pathway | 116 |
| miR-21 | Promotes osteogenic and adipogenic differentiation of hMSCs | 123 |
| miR-24, 125b, 138 | Enhance osteogenic differentiation | 124 |
| miR-27 | miR-27 expression increases during differentiation of human osteoblasts and enhances Wnt signaling by repressing APC | 46 |
| miR-29b, miR-2861 | Increases osteoblast differentiation in pre-osteoblasts by repressing inhibitors of osteogenesis. Promotes BMP2 induced ST2 osteoblast differentiation by repressing histone deacetylase 5 (HDAC5) expression | 113 |
| miR-208 | Enhances BMP2 induced pre-osteoblast differentiation by targeting transactivation of osteopontin and Runx2 | 117 |
| miR-29c | Promotes Wnt signaling in osteoblasts by targeting inhibitors | 115 |
| miR-196a | Overexpression in adipose derived MSCs promotes osteoblast differentiation | 125 |
| hsa-miR-148b | Up-regulate osteoblast differentiation | 126 |
| miR-335-5p | Stimulates Wnt signaling is in mature osteoblasts and regulates Runx2 in MSCs | 116 |
| miR-378 | Promotes osteoblast differentiation by binding with nephronectin | 127 |
| miR-2861/miR-3960 | Repress Runx2 activity in a feed-forward loop | 114 |
| miR-199a, miR-346 | Regulate LIF expression during hMSC differentiation | 128 |
| miR-33a, 204, 211 | Suppressor of osteoclast function | 129,130 |
| miR-10b, miR-218 | Activator of bone metastases development | 131,132 |
| miRNAs:miR-9 | Down regulates muscle transcription factors and thus promotes osteogenesis | 110 |
| miR-191, 449a, 491, 365, 95 and miR-425 | Involves in NF-κB regulation to promote osteoblast differentiation | 133 |
| 11 miRs | Overexpression inhibits/delays osteoblast differentiation | 110 |
| miR-326 | Activator of osteoclast function | 134 |
| miR-29a,b | Attenuates collagen synthesis in mineralized bone | 113 |
| miR-637 | Promotes adipocyte and inhibits osteoblast differentiation via osterix | 116 |
| miR-23a-27a-24-2, 30c, 34c, 133a, 135a, 137, 204, 205, 217, 338 | Decrease expression of Runx2 protein resulting inhibition of osteoblast differentiation | 135 |
| miR-100 | Inhibits osteoblast differentiation in hACSs | 136 |
| miR-133, 135, 138 | Inhibit differentiation of osteoprogenitors through attenuating Runx2 and Smad5 pathways | 118 |
| miR-141 and 200a | Down regulate BMP-2-induced pre-osteoblast differentiation through the translational repression of Dlx5 | 137 |
| miR-206 | Overexpression in osteoblast Inhibits their differentiation | 138 |
On the other hand, there are several miRNAs acting as negative regulators of osteoblast differentiation and mineralization in mesenchymal progenitor cells and bone marrow stromal cell (BMSCs). Runx2 gene expression is inhibited by miR-204 and miR-211.139 MiR-355 is found to be directly targeting Runx2 mRNA, and overexpression of miR-335 in hMSCs inhibits their proliferation and migration, as well as their osteogenic potential.140 TGF-β/BMP signaling pathways and their components in osteoblasts can be inhibited by miRNAs such as hsa-miR-31, hsa-miR-106a, hsa-miR-148a, and hsa-miR-424 and their target genes were found to be as Runx2, Cbfb, BMPs. mir125b and miR26a inhibits osteoblast differentiation by reducing the cell proliferation rate and interference on Smad1 translation, respectively. Yin et al.141 have shown that miR-155 targets the BMP signaling cascade, including Smad1, Smad5, CEBPB, and Runx2. The expression of miR-141 and miR-200a thought to downregulate BMP2-induced pre-osteoblast differentiation through the translational repression of Dlx. Recently, it has been reported that Smads participate in miRNA biogenesis.142 However, these physiological regulation brought out by miRNAs could be important for balancing bone formation and bone resorption processes.
MAPKs and PI3K/AKT pathway and TGF-β/BMP signaling
MAPKs are serine/threonine/tyrosine-specific protein kinases belonging to CMGC (CDK/MAPK/GSK3/CLK) kinase group. Multiple extracellular stimuli can kick off the serial phosphorylation from MAP kinase kinase kinase (MAPKKK) to MAP Kinase.143 However, the linker region of Smad protein is the platform for integrating MAPK/RTK signal with the TGF-β/BMP pathway. The serine rich linker region is loosely organized and structurally flexible, making it accessible for kinases. This region is also rich in threonine and proline residues, favoring phosphorylation by proline directed kinases such as MAPKs.144 Multiple Ser/Thr residues in Smad1 linker can be sequentially phosphorylated by Erk and GSK3-β, creating a docking site for the Smad1/5 specific E3 ubiquitin ligase causes not only ubiquitination and degradation of the Smad but also occludes their interaction with the nuclear pore complex, thereby preventing Smad nuclear translocation and attenuation of the BMP signal.145 But, Wnt/β-catenin signaling, which is known to inactivate GSK3-β, reduces Smad1 ubiquitination and stabilizes the protein.101 More recently, three residues in the linker region of Smad3 (Thr178, Ser203, and Ser207) were identified as Erk1/2 phosphorylation sites both in vitro and in vivo. Erk-mediated phosphorylation of these sites inhibits Smad3 transcriptional activity but does not prevent Smad3 from entering the nucleus146 suggesting the existence of an unknown mechanism for Smad3 inhibition by the linker phosphorylation. Figure 5a shows the role of MAPKs/TAK1 signaling in MSCs differentiation and bone formation.
Studies in Xenopus laevis have indicated that BMPR activation leads to MAPK activation via X-linked inhibitor of apoptosis, TGF-β-activated kinase and Tak-binding protein.147 BMP7 has been shown to induce p38 MAPK through integrin-linked kinases.148 Furthermore, the ligands BMP2 and BMP4 can activate p38 and ERK MAPK, but not JNK. In early mouse embryos, limb-bud outgrowth is promoted by Shh and FGF, whereas termination of this growth requires BMP-mediated inhibition of FGF. On the other hand, defects in cartilage development thought partly because of an elevation in FGF signaling that suppresses chondrogenesis. As a result, BMP function can be suppressed by several signals that activate RTK/MAPK, including EGF, FGF and insulin like growth factor (IGF).149 In addition to R-Smad, MAPKs also phosphorylate and regulate the C-Smad, Smad4, and the inhibitory Smad implicated in the transcriptional regulation of Smad7, therefore indirectly regulating TGF-β signaling.
Turning the focus to how the synergy between the TGF-β and HER2/Ras/MAPK pathways often leads to the secretion of growth factors and cytokines, including TGF-β itself, which in turn promote epithelial-to-mesenchymal transition (EMT). MEK/Erk has been reported to positively regulate Smad3 gene transcription in epithelial cells. Besides MAPK, it has been reported that BMP2 can activate the protein kinase C and PI3K pathways.150 PI3K-activated, plasma membrane-anchored Akt can physically sequester Smad3 and block its nuclear translocation in a kinase-independent manner, without affecting the C-terminal phosphorylation (activation) of Smad3. However, this mechanism may be cell type-specific and dependent on the stoichiometry between the Akt and Smad3 proteins, as Akt can also facilitate Smad3 function, and PI3K/Akt has been shown to be required for the nuclear accumulation of BMP-activated Smad1.150 A third way for PI3K/Akt to debilitate Smad3 is through the inactivation of certain nuclear factors that are necessary for Smad3 function. The regulation of MAPKs by BMP stimulation151 represents an important mechanism for non-Smad TGF-β signaling but the exact mechanism of activation is not so clear.
Notch signaling pathway and TGF-β/BMP signaling
In the nucleus, Notch ICD (intracellular domain) binds specific transcription factors and recruits transcriptional co-activators to induce the expression of bHLH family of genes.152 Notch pathway is active in the early stages of osteoblast differentiation, also by acting on Runx2-dependent osteogenic gene expression.153 Notch could also regulate osteoclastogenesis, through the up-regulation of RANKL and OPG genes, suggesting that the functional cross-talk between osteoblasts and osteoclasts might be also mediated by Notch signaling.152 Notch signaling is also involved in skeletal patterning and somitogenesis.147 Notch1 null mouse embryos exhibits significantly delayed and disorganized somitogenesis.154 Finally, Notch signaling is believed to act also in chondrogenic differentiation, although its exact role and its temporal effects during chondro/osteoblastogenesis are still unclear. Figure 5d summarizes Notch signaling in MSC differentiation. More lately, a sign of the existence of molecular crosstalk between endothelial and osteoblastic cells, angiogenesis in the skeletal system and osteogenesis identified to be coupled, in which Notch signaling is involved. Ramasamy team found that endothelial-cell-specific and inducible genetic disruption of Notch signaling in mice not only impaired bone vessel morphology and growth, but also led to reduced osteogenesis, shortening of long bones, chondrocyte defects, loss of trabeculae and decreased bone mass.155
Other modulators of TGF-β/BMP signaling
There are a number of some other modulators of BMP signaling namely Glucocorticoids (GCs), retinoic acid, laminin, and nitric oxide (NO), nuclear factor kappa B (NF-κB) and hypoxia-inducible factor (HIF). GCs mediate their action on osteoblasts through BMP.156 Retinoic acid is a possible modulator of BMP expression.157 Cell adhesion molecules such as laminin, neural cell adhesion molecules (N-CAM), and integrin are known to interact with BMP and have been localized at the areas of initial mesenchymal condensation158,159. TGF-β activities are also interconnected with mTOR and NO signaling. Lamouille et al.160 reported that mTORC2 is a novel and crucial mediator in TGF-β signaling. Upon TGF-β activation, Smad controls the expression and activities of Snai1 transcription factor involved in EMT, which represses transcription of the E-cadherin gene.161 TGF-β-induced Akt (S473) phosphorylation is dependent on mTORC2 activity.160 Akt blocks phospho-activation of Smad3 by an Akt kinase-dependent mechanism through mTOR and also blocks TGF-β signals downstream of Smad3 activation, but through a mechanism that does not require the kinase activity of Akt or mTOR.162 Attenuated NO and small interfering RNA based silencing of secreted modular calcium-binding protein 1 (SMOC-1) decreased TGF-β, reduced Smad binding to DNA, and decreased mRNA expression of genes regulated by TGF-β.163 In particular, NO reduced the metabolic life of ectopically expressed Smad2 and enhanced its ubiquitination. The endothelial NO/cGMP/PKG pathway interferes with TGF-β/Smad2 signaling by directing the proteasomal degradation of activated Smad.164 Ubiquitously expressed NF-κB transcription factors and HIF also recently found to be involved in TGF-β/BMP signaling. HIF-2α regulates collagen X, MMP-13 and VEGF expression.165 In addition, further possible transcription targets of HIF-2α relates to endochondral ossification have been identified, namely Runx2 and Ihh proteins. Concerning activated NF-κB that influences the amount and remodeling of ECM proteins, and shows indirect positive effects on downstream regulators including HIF-2α, β-catenin and Runx2.166 Concerning bone, NF-κB proteins induce the synthesis of BMP2. In normal conditions RANK/RANKL interaction is regulated by the OPG decoy receptor that by blocking RANKL promotes bone formation.167 Contrary, the interaction between the IL-1/TNF induced RANKL and RANK surface receptor activates NF-κB turn out PGE2 and thereby bone resorption.168
Sometimes BMP cannot generate enough clinical response to bone regeneration treatment. A possible reason might be that inflammatory cytokines inhibit the bone formation and osteoblast differentiation induced by BMP. For example, one inflammatory cytokine, tumor necrosis factor (TNF) a, inhibits osteoblast differentiation in multiple models, including fetal calvaria, bone marrow stromal cells. Together, there seem to be antagonistic effects between BMP/Smad and NF-κB signaling systems. Although the involvement of NF-κB in osteoclastogenesis has been well investigated, little is currently known about the possible involvement of NF-κB in osteoblast differentiation, especially the cross-talk between BMP/Smad and NF-κB signaling in osteoblast differentiation.169 NF-κB activation inhibits osteoblast differentiation at least in part by interfering with Smad signaling. NF-κB is involved in the inhibitory effect of TNFa on BMP2-induced osteoblast differentiation. Eliseev et al. reported that Inhibition of NF-κB represses BMP/Smad signaling and BMP2-induced differentiation through Smad7 leads to the induction of osteoblast differentiation.170 Inhibition of NF-κB could suppress the antiosteoblastogenic activity of TNFa and relieve the suppression of BMP2-induced Runx2 expression mediated by TNFa.171 Hence, TNFa on BMP signaling in osteoblast differentiation is closely linked to the NF-κB pathway. NF-κB activation inhibits osteoblast differentiation at least in part by interfering with Smad signaling. An intracellular balance of signal intensities between NF-κB and BMP/Smad is therefore crucial for osteoblast differentiation.
TGF-β/BMP signaling are finely tuned by a complex network of signaling molecules and biophysical factors, shedding light on some extremely relevant modulators namely FGF, HIF, GC, NO, PI3K, Akt, NF-κB, and mTOR. On a final note it could be remarked that given the considerable variation in the use of receptors, abundance of transcriptional co-regulators and links to other major pathways TGF-β/BMP signaling creates more of a signaling network than a straight linear regulatory path.
Bone microenvironment and bone formation
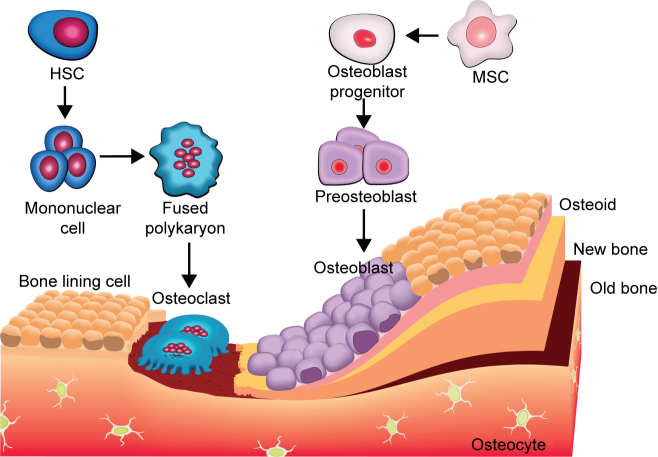
MSCs derived osteoblasts, osteoclasts, mineralized bone matrix, and some sort of fixed osteocytes within bone are the major components of bone microenvironment.40 Formation of new bone and the regulation of osteoclastogenesis are two main functions of the osteoblasts.172 Osteogenesis comprises a sequential cascade with three critical phases: migration and mitosis of mesenchymal cells, differentiation of mesenchymal cells into chondroblasts, cartilage formation and, finally, substitution of cartilage by bone. During osteogenesis some pluripotent mesenchymal cell lines, bone marrow cells, and osteoblast precursors are the target of BMP. The pluripotent stem cells undergo successive stages of differentiation with a decreasing proliferation potential, giving rise to committed pre-osteoblasts. Subsequently, pre-osteoblasts differentiate into mature osteoblasts that deposit the necessary components to form bone matrix, followed by mineralization.173 Eventually, mature, mineralizing osteoblasts become embedded in the newly secreted bone matrix and undergo terminal differentiation to form osteocytes. Figure 6 shows the evolution of osteoblasts and osteoclasts in bone formation.
Osteoclasts are large multinucleated terminally differentiated cells with a unique ability for bone resorption. They are derived from hematopoietic stem cells. The cells undergo proliferation in response to M-CSF. The precursor cells flaunt receptor and activator of nuclear factor κB (RANK) on the surface, while the ligand RANKL is expressed by the bone marrow stromal cells and osteoblasts.174 Binding of the ligand to the receptor commits the precursor cells to differentiate into osteoclast lineage. The same interaction is also critical for osteoclast formation and can also promote osteoclast activity, since RANK is also present on the surface of terminally differentiated osteoclasts. Osteoprotegerin (OPG) is a soluble decoy receptor and a competitor of RANKL in its binding with RANK and thus can inhibit osteoclastogenesis.175 Therefore, the balance of RANKL and OPG is critical for osteoclast formation and activity. Cathepsin K and MMPs catalyze the degradation of bone matrix, dissolution of the bone mineral, and resorption of the bone.176 A recent study on OVX mice treated with exogenous PDGF-BB or inhibition of cathepsin K showed that, osteogenesis during bone modeling and remodeling is coupled with angiogenesis.177
In general active BMP dimers act as a chemo attractant for undifferentiated mesenchymal stem cells, causing them to move into the area of bone defect or injury and over expression of activated BMPRs ultimately increased production of bone matrix proteins.178 Receptor bounded BMP is displayed on the stem cells' surface. The Smad complex translocates into the nucleus and interacts with several transcription factors such as Runx2/Cbfα1, Osx, Dlx5, and Msx2. The expression of osteoblast genes such as alkaline phosphatase, osteopontin (Opn), and bone sialoprotein are also found to be important for osteogenesis. For instance, activated Smad relieves repression of the osteopontin gene by binding to the repressor Hoxc8 and removing it from the Opn promoter. Hoxc8, a repressor in the BMP pathway binds to the consensus site of the Opn gene and silences its transcription. The Smad1/4 complex, in response to BMP, interacts with its DNA-binding domain and dislodges it from the OPN promotor element, thereby initiating gene transcription and inducing osteoblast differentiation.179 Recently in mouse model it is shown that notch signaling pathway enhances BMP2 responsiveness of Msx2 gene to induce osteogenic differentiation.180 In addition to others, menin is required for BMP-induced osteoblast differentiation.181 Nonetheless, BMP2 recruit's mesenchymal cells surrounding the initial cartilage condensations into chondrogenic fate, BMP6 accelerates hypertrophic and terminal chondrocyte differentiation and mineral accretion while GDF5 is required for joint formation.179 In multipotential mesenchymal cells isolated from human bone marrow, BMP2 and 9 promote chondrogenic differentiation, possibly through activation of Sox-9, a chondrogenic-related transcription factor.182 It is frequently assumed that BMP-activated Smads directly regulate the expression of Runx2 and members of the BMP family of growth/differentiation factors. But the interactions between Runx2 and BMP-specific Smads (Smad1 and 5) suggest their intrinsic cooperation in mediating BMP functions. Although the mechanism is not fully clear. It has been proposed that BMPs can induce both undifferentiated stem cells and more differentiated multipotent cells along chondrogenic or osteogenic pathways. Figure 5 summarizes the up to date integration of different signaling pathways for lineage specific differentiation for MSCs in an exquisitely coordinated manner. Evidently behind its outward rigidity, bone is a highly dynamic organ where homeostasis is tightly controlled and largely dependent upon cellular communication between osteoclasts and osteoblasts. This tight coupling between bone resorption and bone formation is essential for the correct function and maintenance of the skeletal system, repairing microscopic skeletal damage, and replacing aged bone.
Concluding remarks
To date, a large number of regulatory factors and targets have been identified for the TGF-β/BMP signaling pathway, leading to a complex interactive network that investigators are still trying to unravel. High throughput screening, modeling, structural biology, and bioinformatics will greatly assist the elucidation of the complex mechanisms regarding TGF-β/BMP function and regulation to close the gaps. Knowing how these pathways affect the transformation of osteoblasts to osteocytes in different developmental stages help to exploit the therapy of bone related diseases. However, BMP was named for the ability to form bone, and the name implies that they are capable of inducing the osteogenic differentiation of non-bone cells. This is true for most of the best-studied BMPs, but not all. Furthermore, the name does not acknowledge the breadth of BMP actions, and it is still not clear what controls and how a specific BMP will be osteoinductive. And the number of players in osteogenesis is much wider than that described in this review, and much remains to be understood about the complexity of BMP signaling.
Acknowledgments
We are grateful to Prof. Pritinder Kaur (Epithelial Stem Cell Biology Laboratory, Peter MacCallum Cancer Centre, Australia) for her helpful suggestion and assistance. We also gratefully thank S.M. Abdul Awal, Signal Transduction Group, Cambridge University, UK, for fruitful discussions.
Notes
The authors declare no conflict of interest.
References
- Beederman M, Lamplot JD, Nan G. BMP signaling in mesenchymal stem cell differentiation and bone formation. J Biomed Sci Eng. 2013;6:32–52. [Europe PMC free article] [Abstract] [Google Scholar]
- Botchkarev VA. Bone morphogenetic proteins and their antagonists in skin and hair follicle biology. J Invest Dermatol. 2003;120:36–47. [Abstract] [Google Scholar]
- Lechleider RJ, Ryan JL, Garrett L. Targeted mutagenesis of Smad1 reveals an essential role in chorioallantoic fusion. Dev Biol. 2001;240:157–167. [Abstract] [Google Scholar]
- Zhao GQ. Consequences of knocking out BMP signaling in the mouse. Genesis. 2003;35:43–56. [Abstract] [Google Scholar]
- Urist MR. Bone morphogenetic protein: the molecularization of skeletal system development. J Bone Miner Res. 1997;12:343–346. [Abstract] [Google Scholar]
- Carreira AC, Lojudice FH, Halcsik E, Navarro RD, Sogayar MC, Granjeiro JM. Bone morphogenetic proteins: facts, challenges, and future perspectives. J Dent Res. 2014;93:335–345. [Abstract] [Google Scholar]
- Feng XH, Derynck R. Specificity and versatility in TGF-β signaling through Smads. Annu Rev Cell Dev Biol. 2005;21:659–693. [Abstract] [Google Scholar]
- Balemans W, Hul WV. Extracellular regulation of BMP signaling in vertebrates: a cocktail of modulators. Dev Biol. 2002;250:231–250. [Abstract] [Google Scholar]
- Moustakas A, Heldin CH. The regulation of TGF-β signal transduction. Development. 2009;136:3699–3714. [Abstract] [Google Scholar]
- Schmierer B, Hill CS. TGF-β-SMAD signal transduction: molecular specificity and functional flexibility. Nat Rev Mol Cell Biol. 2007;8:970–982. [Abstract] [Google Scholar]
- Guo L, Zhao RC, Wu Y. The role of microRNAs in self-renewal and differentiation of mesenchymal stem cells. Exp Hematol. 2011;39:608–616. [Abstract] [Google Scholar]
- Bessa PC, Casal M, Reis RL. Bone morphogenetic proteins in tissue engineering: the road from the laboratory to the clinic, part I (basic concepts) J Tissue Eng Regen Med. 2008;2:1–13. [Abstract] [Google Scholar]
- Pan A, Chang L, Nguyen A, James AW. A review of hedgehog signaling in cranial bone development. Front Physiol. 2013;4:61. [Europe PMC free article] [Abstract] [Google Scholar]
- Horikiri Y, Shimo T, Kurio N. Sonic hedgehog regulates osteoblast function by focal adhesion kinase signaling in the process of fracture healing. PLoS ONE. 2013;8:e76785. [Europe PMC free article] [Abstract] [Google Scholar]
- Reichert JC, Schmalzl J, Prager P. Synergistic effect of Indian hedgehog and bone morphogenetic protein-2 gene transfer to increase the osteogenic potential of human mesenchymal stem cells. Stem Cell Res Ther. 2013;4:105. [Europe PMC free article] [Abstract] [Google Scholar]
- Kim JH, Liu X, Wang J. Wnt signaling in bone formation and its therapeutic potential for bone diseases. Ther Adv Musculoskelet Dis. 2013;5:13–31. [Europe PMC free article] [Abstract] [Google Scholar]
- Issack PS, Helfet DL, Lane JM. Role of Wnt signaling in bone remodeling and repair. HSS J. 2008;4:66–70. [Europe PMC free article] [Abstract] [Google Scholar]
- Terada K, Misao S, Katase N, Nishimatsu S, Nohno T. Interaction of Wnt Signaling with BMP/Smad Signaling during the Transition from Cell Proliferation to Myogenic Differentiation in Mouse Myoblast-Derived Cells. Int J Cell Biol. 2013;2013:616294. [Europe PMC free article] [Abstract] [Google Scholar]
- Lin GL, Hankenson KD. Integration of BMP, Wnt, and notch signaling pathways in osteoblast differentiation. J Cell Biochem. 2011;112:3491–3501. [Europe PMC free article] [Abstract] [Google Scholar]
- Balboni AL, Hutchinson JA, DeCastro AJ. ΔNp63α mediated activation of bone morphogenetic protein signaling governs stem cell activity and plasticity in normal and malignant mammary epithelial cells. Cancer Res. 2013;73:1020–1030. [Europe PMC free article] [Abstract] [Google Scholar]
- Cheng H, Jiang W, Phillips FM. Osteogenic activity of the fourteen types of human bone morphogenetic proteins (BMPs) J Bone Joint Surg Am. 2003;85-A:1544–1552. [Abstract] [Google Scholar]
- Kang Q, Sun MH, Cheng H. Characterization of the distinct orthotopic bone-forming activity of 14 BMPs using recombinant adenovirus-mediated gene delivery. Gene Therapy. 2004;11:1312–1320. [Abstract] [Google Scholar]
- Luther G, Wagner ER, Zhu G. BMP-9 induced osteogenic differentiation of mesenchymal stem cells: molecular mechanism and therapeutic potential. Curr Gene Ther. 2011;11:229–240. [Abstract] [Google Scholar]
- Lamplot JD, Qin J, Nan G. BMP9 signaling in stem cell differentiation and osteogenesis. Am J Stem Cells. 2013;2:1–21. [Europe PMC free article] [Abstract] [Google Scholar]
- Rivera JC, Strohbach CA, Wenke JC, Rathbone CR. Beyond osteogenesis: an in vitro comparison of the potentials of six bone morphogenetic proteins. Front Pharmacol. 2013;4:125. [Europe PMC free article] [Abstract] [Google Scholar]
- Callis TE, Cao D, Wang DZ. Bone morphogenetic protein signaling modulates myocardin transactivation of cardiac genes. Circ Res. 2005;97:992–1000. [Europe PMC free article] [Abstract] [Google Scholar]
- Samartzis D, Khanna N, Shen FH, An HS. Update on bone morphogenetic proteins and their application in spine surgery. J Am Coll Surg. 2005;200:236–248. [Abstract] [Google Scholar]
- Gamer LW, Cox K, Carlo JM, Rosen V. Overexpression of BMP3 in the developing skeleton alters endochondral bone formation resulting in spontaneous rib fractures. Dev Dyn. 2009;238:2374–2381. [Europe PMC free article] [Abstract] [Google Scholar]
- Knöchel S, Dillinger K, Köster M, Knöchel W. Structure and expression of Xenopus tropicalis BMP-2 and BMP-4 genes. Mech Dev. 2001;109:79–82. [Abstract] [Google Scholar]
- Zuzarte-Luís V, Montero JA, Rodriguez-León J, Merino R, Rodríguez-Rey JC, Hurlé JM. A new role for BMP5 during limb development acting through the synergic activation of Smad and MAPK pathways. Dev Biol. 2004;272:39–52. [Abstract] [Google Scholar]
- Paralkar VM, Grgurevic L, Simic P. A novel role of bone morphogenetic protein-6 (BMP-6) as an endocrine regulator of bone and glucose homeostasis. Bone. 2010;47:S57. [Google Scholar]
- Tsumaki N, Yoshikawa H. The role of bone morphogenetic proteins in endochondral bone formation. Cytokine Growth Factor Rev. 2005;16:279–285. [Abstract] [Google Scholar]
- Lopez-Coviella I, Mellott TM, Kovacheva VP. Developmental pattern of expression of BMP receptors and Smads and activation of Smad1 and Smad5 by BMP9 in mouse basal forebrain. Brain Res. 2006;1088:49–56. [Abstract] [Google Scholar]
- Andersson O, Reissmann E, Ibanez CF. Growth differentiation factor 11 signals through the transforming growth factor-beta receptor ALK5 to regionalize the anterior–posterior axis. EMBO Rep. 2006;7:831–837. [Europe PMC free article] [Abstract] [Google Scholar]
- Shen B, Bhargav D, Wei A. BMP-13 emerges as a potential inhibitor of bone formation. Int J Biol Sci. 2009;5:192–200. [Europe PMC free article] [Abstract] [Google Scholar]
- Di Pasquale E, Beck-Peccoz P, Persani L. Hypergonadotropic ovarian failure associated with an inherited mutation of human bone morphogenetic protein-15 (BMP15) gene. Am J Hum Genet. 2004;75:106–111. [Europe PMC free article] [Abstract] [Google Scholar]
- Nishimoto Y, Morisaki H, Yamada O, Ichinose Y, Suzuki N. Japanese case of hereditary hemorrhagic telangiectasia type 2 with a novel mutation, c.154A>C (p.Thr52Pro), in the ALK1/ACVRL1 gene. Neurol Clin Neurosci. 2014;2:126–128. [Google Scholar]
- Fukuda T, Kohda M, Kanomata K. Constitutively activated ALK2 and increased SMAD1/5 cooperatively induce bone morphogenetic protein signaling in fibrodysplasia ossificans progressive. J Biol Chem. 2009;284:7149–7156. [Europe PMC free article] [Abstract] [Google Scholar]
- Howe JR, Bair JL, Sayed MG. Germ line mutations of the gene encoding bone morphogenetic protein receptor 1A in juvenile polyposis. Nat Genet. 2001;28:184–187. [Abstract] [Google Scholar]
- Chen G, Deng C, Li YP. TGF-β and BMP signaling in osteoblast differentiation and bone formation. Int J Biol Sci. 2012;8:272–288. [Europe PMC free article] [Abstract] [Google Scholar]
- Fessel JP, Loyd JE, Austin ED. The genetics of pulmonary arterial hypertension in the post-BMPR2 era. Pulm Circ. 2011;1:305–319. [Europe PMC free article] [Abstract] [Google Scholar]
- Pearsall RS, Canalis E, Cornwall-Brady M. A soluble activin type IIA receptor induces bone formation and improves skeletal integrity. Proc Natl Acad Sci U S A. 2008;105:7082–7087. [Europe PMC free article] [Abstract] [Google Scholar]
- Goncalves MD.The role of the activin receptor type IIb (ActRIIB) in glucose and energy homeostasis . Dissertations available from ProQuest 2010; Paper AAI3447175. http://repository.upenn.edu/dissertations/AAI3447175
- Matsunobu T, Torigoe K, Ishikawa M. Critical roles of the TGF-β type I receptor ALK5 in perichondrial formation and function, cartilage integrity, and osteoblast differentiation during growth plate development. Dev Biol. 2009;332:325–338. [Europe PMC free article] [Abstract] [Google Scholar]
- Miyazono K, Kamiya Y, Morikawa M. Bone morphogenetic protein receptors and signal transduction. J Biochem. 2010;147:35–51. [Abstract] [Google Scholar]
- Wang T, Xu Z. miR-27 promotes osteoblast differentiation by modulating Wnt signaling. Biochem Biophys Res Commun. 2010;402:186–189. [Abstract] [Google Scholar]
- Massagué J, Seoane J, Wotton D. Smad transcription factors. Genes Dev. 2005;19:2783–2810. [Abstract] [Google Scholar]
- Estrada KD, Retting KN, Chin AM, Lyons KM. Smad6 is essential to limit BMP signaling during cartilage development. J Bone Miner Res. 2011;26:2498–2510. [Europe PMC free article] [Abstract] [Google Scholar]
- Song Q, Zhu B, Hu W. A common SMAD7 variant is associated with risk of colorectal cancer: evidence from a case-control study and a meta-analysis. PLoS ONE. 2012;7:e33318. [Europe PMC free article] [Abstract] [Google Scholar]
- Okuda T, van Deursen J, Hiebert SW, Grosveld G, Downing JR. AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell. 1996;84:321–330. [Abstract] [Google Scholar]
- Wu CJ, Lu HK. Smad signal pathway in BMP-2-induced osteogenesis a mini review. J Dent Sci. 2008;3:13–21. [Google Scholar]
- Inoue K, Ozaki S, Shiga T. Runx3 controls the axonal projection of proprioceptive dorsal root ganglion neurons. Nat Neurosci. 2002;5:946–954. [Abstract] [Google Scholar]
- Constam DB, Robertson EJ. Regulation of bone morphogenetic protein activity by prodomains and proprotein convertases. J Cell Biol. 1999;144:139–149. [Europe PMC free article] [Abstract] [Google Scholar]
- Szpalski M, Gunzburg R. Recombinant human bone morphogenetic protein-2: a novel osteoinductive alternative to autogenous bone graft? Acta Orthop Belg. 2005;71:133–148. [Abstract] [Google Scholar]
- Arrabal PM, Visser R, Santos-Ruiz L, Becerra J, Cifuentes M. Osteogenic molecules for clinical applications: improving the BMP-collagen system. Biol Res. 2013;46:421–429. [Abstract] [Google Scholar]
- Granjeiro JM, Oliveira RC, Bustos-Valenzuela JC, Sogayar MC, Taga R. Bone morphogenetic proteins: from structure to clinical use. Braz J Med Biol Res. 2005;38:1463–1473. [Abstract] [Google Scholar]
- Rao SM, Ugale GM, Warad SB. Bone morphogenetic proteins: periodontal regeneration. N Am J Med Sci. 2013;5:161–168. [Europe PMC free article] [Abstract] [Google Scholar]
- Muenster U, Korupolu R, Rastogi R, Read J, Fischer WH. Antagonism of activin by activin chimeras. Vitam Horm. 2011;85:105–128. [Europe PMC free article] [Abstract] [Google Scholar]
- Samad TA, Rebbapragada A, Bell E. DRAGON, a bone morphogenetic protein co-receptor. J Biol Chem. 2005;280:14122–14129. [Abstract] [Google Scholar]
- Bonor J, Adams EL, Bragdon B, Moseychuk O, Czymmek KJ, Nohe A. Initiation of BMP2 signaling in domains on the plasma membrane. J Cell Physiol. 2012;227:2880–2888. [Europe PMC free article] [Abstract] [Google Scholar]
- Allendorph GP, , Vale WW, Choe S. Structure of the ternary signaling complex of a TGF-beta superfamily member. Proc Natl Acad Sci U S A. 2006;103:7643–7648. [Europe PMC free article] [Abstract] [Google Scholar]
- Yu PB, Beppu H, Kawai N, Li E, Bloch KD. Bone morphogenetic protein (BMP) type II receptor deletion reveals BMP ligand-specific gain of signaling in pulmonary artery smooth muscle cells. J Biol Chem. 2005;280:24443–24450. [Abstract] [Google Scholar]
- Miyazono K, Maeda S, Imamur T. BMP receptor signaling: transcriptional targets, regulation of signals, and signaling cross-talk. Cytokine Growth Factor Rev. 2005;16:251–263. [Abstract] [Google Scholar]
- Shore EM, Xu M, Feldman GJ. A recurrent mutation in the BMP type I receptor ACVR1 causes inherited and sporadic fibrodysplasia ossificans progressive. Nat Genet. 2006;38:525–527. [Abstract] [Google Scholar]
- Morovvati Z, SMorovvati S, Alishiri GA, Moosavi SH, Ranjbar R, Moghaddam YB. Mutation detection in activin A receptor, Type I (ACVR1) gene in fibrodysplasia ossificans progressiva in an Iranian family. Cell J. 2014;16:91–94. [Europe PMC free article] [Abstract] [Google Scholar]
- Chalmers J, Gray DH, Rush J. Observations on the induction of bone in soft tissues. J Bone Joint Surg Br. 1975;57:36–45. [Abstract] [Google Scholar]
- Kaplan FS, Le Merrer M, Glaser DL. Fibrodysplasia ossificans progressiva. Best Pract Res Clin Rheumatol. 2008;22:191–205. [Europe PMC free article] [Abstract] [Google Scholar]
- Kaplan FS, Groppe J, Shore EM. When one skeleton is enough: approaches and strategies for the treatment of fibrodysplasia ossificans progressiva (FOP) Drug Discov Today Ther Strateg. 2008;5:255–262. [Europe PMC free article] [Abstract] [Google Scholar]
- Yu PB, Deng DY, Lai CS. BMP type I receptor inhibition reduces heterotopic ossification. Nat Med. 2008;14:1363–1369. [Europe PMC free article] [Abstract] [Google Scholar]
- Gregson CL, Hollingworth P, Williams M. A novel ACVR1 mutation in the glycine/serine-rich domain found in the most benign case of a fibrodysplasia ossificans progressiva variant reported to date. Bone. 2011;48:654–658. [Europe PMC free article] [Abstract] [Google Scholar]
- Kaplan FS, Xu M, Seemann P. Classic and atypical fibrodysplasia ossificans progressiva (FOP) phenotypes are caused by mutations in the bone morphogenetic protein (BMP) type I receptor ACVR1. Hum Mutat. 2009;30:379–390. [Europe PMC free article] [Abstract] [Google Scholar]
- Petrie KA, Lee WH, Bullock AN. Novel mutations in ACVR1 result in atypical features in two fibrodysplasia ossificans progressiva patients. PLoS ONE. 2009;4:e5005. [Europe PMC free article] [Abstract] [Google Scholar]
- Ratbi I, Borcciadi R, Regragui A, Ravazzolo R, Sefiani A. Rarely occurring mutation of ACVR1 gene in Moroccan patient with fibrodysplasia ossificans progressiva. Clin Rheumatol. 2010;29:119–121. [Abstract] [Google Scholar]
- Carvalho DR, Navarro MM, Martins BJ. Mutational screening of ACVR1 gene in Brazilian fibrodysplasia ossificans progressiva patients. Clin Genet. 2010;77:171–176. [Abstract] [Google Scholar]
- Furuya H, Ikezoe K, Wang L. A unique case of fibrodysplasia ossificans progressiva with an ACVR1 mutation, G356D, other than the common mutation (R206H) Am J Med Genet A. 2008;146A:459–463. [Abstract] [Google Scholar]
- Bocciardi R, Bordo D, Di Duca M, Di Rocco M, Ravazzolo R. Mutational analysis of the ACVR1 gene in Italian patients affected with fibrodysplasia ossificans progressiva: confirmations and advancements. Eur J Hum Genet. 2009;17:311–318. [Europe PMC free article] [Abstract] [Google Scholar]
- Chaikuad A, Alfano I, Kerr G. Structure of the bone morphogenetic protein receptor ALK2 and implications for fibrodysplasia ossificans progressiva. J Biol Chem. 2012;287:36990–36998. [Europe PMC free article] [Abstract] [Google Scholar]
- Bagarova J, Vonner AJ, Armstrong KA. Constitutively active ALK2 receptor mutants require Type II receptor cooperation. Mol Cell Biol. 2013;33:2413–2424. [Europe PMC free article] [Abstract] [Google Scholar]
- Lowery JW, Rosen V. Allele-specific RNA interference in FOP silencing the FOP gene. Gene Therapy. 2012;19:701–702. [Abstract] [Google Scholar]
- Wu JW, Hu M, Chai J. Crystal structure of a phosphorylated Smad2. Recognition of phosphoserine by the MH2 domain and insights on Smad function in TGF-beta signaling. Mol Cell. 2001;8:1277–1289. [Abstract] [Google Scholar]
- Makkar P, Metpally RP, Sangadala S, Reddy BV. Modeling and analysis of MH1 domain of Smads and their interaction with promoter DNA sequence motif. J Mol Graph Model. 2009;27:803–812. [Abstract] [Google Scholar]
- ten Dijke P, Miyazono K, Heldin CH. Signaling inputs converge on nuclear effectors in TGF-beta signaling. Trends Biochem Sci. 2000;25:64–70. [Abstract] [Google Scholar]
- Zhang J, Fei T, Li Z, Zhu G, Wang L, Chen Y. BMP induces cochlin expression to facilitate self-renewal and suppress neural differentiation of mouse embryonic stem cells. J Biol Chem. 2013;288:8053–8060. [Europe PMC free article] [Abstract] [Google Scholar]
- Lo RS, Chen YG, Shi Y, Pavletich NP, Massague J. The L3 loop: a structural motif determining specific interactions between SMAD proteins and TGF-beta receptors. Embo J. 1998;17:996–1005. [Europe PMC free article] [Abstract] [Google Scholar]
- Liu F, Hata A, Baker JC. A human Mad protein acting as a BMP-regulated transcriptional activator. Nature. 1996;381:620–623. [Abstract] [Google Scholar]
- Kusanagi K, Inoue H, Ishidou Y, Mishima HK, Kawabata M, Miyazono K. Characterization of a bone morphogenetic protein-responsive Smad-binding element. Mol Biol Cell. 2000;11:555–565. [Europe PMC free article] [Abstract] [Google Scholar]
- Mukhopadhyay P, Webb CL, Warner DR, Greene RM, Pisano MM. BMP signaling dynamics in embryonic orofacial tissue. J Cell Physiol. 2008;216:771–779. [Europe PMC free article] [Abstract] [Google Scholar]
- Morikawa M, Koinuma D, Tsutsumi S. ChIP-seq reveals cell type-specific binding patterns of BMP-specific Smads and a novel binding motif. Nucleic Acids Res. 2011;39:8712–8727. [Europe PMC free article] [Abstract] [Google Scholar]
- Ross S, Cheung E, Petrakis TG, Howell M, Kraus WL, Hill CS. Smads orchestrate specific histone modifications and chromatin remodeling to activate transcription. EMBO J. 2006;25:4490–4502. [Europe PMC free article] [Abstract] [Google Scholar]
- James AW. Review of Signaling Pathways Governing MSC Osteogenic and Adipogenic Differentiation. Scientifica (Cairo) 2013;2013:684736. [Europe PMC free article] [Abstract] [Google Scholar]
- Zhao M, Qiao M, Harris SE, Chen D, Oyajobi BO, Mundy GR. The zinc finger transcription factor Gli2 mediates bone morphogenetic protein 2 expression in osteoblasts in response to hedgehog signaling. Mol Cell Biol. 2006;26:6197–6208. [Europe PMC free article] [Abstract] [Google Scholar]
- Edwards PC, Ruggiero S, Fantasia J. Sonic hedgehog gene-enhanced tissue engineering for bone regeneration. Gene Therapy. 2005;12:75–86. [Abstract] [Google Scholar]
- Karamboulas K, Dranse HJ, Underhill TM. Regulation of BMP-dependent chondrogenesis in early limb mesenchyme by TGF beta signals. J Cell Sci. 2010;123:2068–2076. [Abstract] [Google Scholar]
- Yuasa T, Kataoka H, Kinto N. Sonic hedgehog is involved in osteoblast differentiation by cooperating with BMP-2. J Cell Physiol. 2002;193:225–232. [Abstract] [Google Scholar]
- Deschaseaux F, Sensebe L, Heymann D. Mechanisms of bone repair and regeneration. Trends Mol Med. 2009;15:417–429. [Abstract] [Google Scholar]
- Zhang J, Tan X, Li W. Smad4 is required for the normal organization of the cartilage growth plate. Dev Biol. 2005;284:311–322. [Abstract] [Google Scholar]
- Kalderon D. Similarities between the Hedgehog and Wnt signaling pathways. Trends Cell Biol. 2002;12:523–531. [Abstract] [Google Scholar]
- Taipale J, Beachy PA. The Hedgehog and Wnt signaling pathways in cancer. Nature. 2001;411:349–354. [Abstract] [Google Scholar]
- Komiya Y, Habas R. Wnt signal transduction pathways. Organogenesis. 2008;4:68–75. [Europe PMC free article] [Abstract] [Google Scholar]
- Cadigan KM, Liu YI. Wnt signaling: complexity at the surface. J Cell Sci. 2006;119:395–402. [Abstract] [Google Scholar]
- Fuentealba LC, Eivers E, Ikeda A. Integrating patterning signals: Wnt/GSK3 regulates the duration of the BMP/Smad1 signal. Cell. 2007;131:980–993. [Europe PMC free article] [Abstract] [Google Scholar]
- Krishnan V, Bryant HU, MacDougald OA. Regulation of bone mass by Wnt signaling. J Clin Invest. 2006;116:1202–1209. [Europe PMC free article] [Abstract] [Google Scholar]
- Okamoto M, Udagawa N, Uehara S. Noncanonical Wnt5a enhances Wnt/b-catenin signaling during Osteoblastogenesis. Sci Rep. 2014;4:4493. [Europe PMC free article] [Abstract] [Google Scholar]
- Jin EJ, Lee SY, Choi YA, Jung JC, Bang OS, Kang SS. BMP-2 enhanced chondrogenesis involves p38 MAPK-mediated downregulation of Wnt-7a pathway. Mol Cells. 2006;22:353–359. [Abstract] [Google Scholar]
- Papathanasiou I, Malizos KN, Tsezou A. Bone morphogenetic protein-2-induced Wnt/β-catenin signaling pathway activation through enhanced low-density-lipoprotein receptor-related protein 5 catabolic activity contributes to hypertrophy in osteoarthritic chondrocytes. Arthritis Res Ther. 2012;14:R82. [Europe PMC free article] [Abstract] [Google Scholar]
- Soltanoff CS, Chen W, Yang S, Li YP. Signaling networks that control the lineage commitment and differentiation of bone cells. Crit Rev Eukaryot Gene Expr. 2009;19:1–46. [Europe PMC free article] [Abstract] [Google Scholar]
- McCarthy TL, Centrella M. Novel links among Wnt and TGF-β signaling and Runx2. Mol Endocrinol. 2010;24:587–597. [Europe PMC free article] [Abstract] [Google Scholar]
- Hill TP, Später D, Taketo MM, Birchmeier W, Hartmann C. Canonical Wnt/β-catenin signaling prevents osteoblasts from differentiating into chondrocytes. Dev Cell. 2005;8:727–738. [Abstract] [Google Scholar]
- Schramke V, Hairpin AR. RNAs and retrotransposon LTRs effect RNAi and chromatin-based gene silencing. Science. 2003;301:1069–1074. [Abstract] [Google Scholar]
- Lian JB, Stein GS, van Wijnen AJ. MicroRNA control of bone formation and homeostasis. Nat Rev Endocrinol. 2012;8:212–227. [Europe PMC free article] [Abstract] [Google Scholar]
- He X, Eberhart JK, Postlethwait JH. MicroRNAs and micromanaging the skeletonin disease, development and evolution. J Cell Mol Med. 2009;13:606–618. [Europe PMC free article] [Abstract] [Google Scholar]
- Jafferji M. Regulation of osteoblast differentiation by microRNAs. Umass Med School. 2009.
- Li Z, Hassan MQ, Jafferji M. Biological functions of miR-29b contribute to positive regulation of osteoblast differentiation. J Biol Chem. 2009;284:15676–15684. [Europe PMC free article] [Abstract] [Google Scholar]
- Hu R, Liu W, Li H. A Runx2/miR-3960/miR-2861 regulatory feedback loop during mouse osteoblast differentiation. J Biol Chem. 2011;286:12328–12339. [Europe PMC free article] [Abstract] [Google Scholar]
- Kapinas K, Kessler C, Ricks T, Gronowicz G, Delany AM. miR-29 modulates Wnt signaling in human osteoblasts through a positive feedback loop. J Biol Chem. 2010;285:25221–25231. [Europe PMC free article] [Abstract] [Google Scholar]
- Zhang J, Tu Q, Bonewald LF. Effects of miR-335-5p in modulating osteogenic differentiation by specifically down regulating Wnt antagonist DKK1. J Bone Miner Res. 2011;26:1953–1963. [Europe PMC free article] [Abstract] [Google Scholar]
- Itoh T, Takeda S, Akao Y. MicroRNA-208 modulates BMP-2-stimulated mouse preosteoblast differentiation by directly targeting V-ets erythroblastosis virus E26 oncogene Homolog 1. J Biol Chem. 2010;285:27745–27752. [Europe PMC free article] [Abstract] [Google Scholar]
- Eskildsen T, Taipaleenmäki H, Stenvang J. MicroRNA-138 regulates osteogenic differentiation of human stromal (mesenchymal) stem cells in vivo. Proc Natl Acad Sci. 2011;108:6139–6144. [Europe PMC free article] [Abstract] [Google Scholar]
- Liu Y, Liu W, Hu C. MiR-17 modulates osteogenic differentiation through a coherent feed-forward loop in mesenchymal stem cells isolated from periodontal ligaments of patients with periodontitis. Stem Cells. 2011;29:1804–1816. [Abstract] [Google Scholar]
- Mizoguchi F, Murakami Y, Saito T, Miyasaka N, Kohsaka H. MiR-31 controls osteoclast formation and bone resorption by targeting RhoA. Arthritis Res Ther. 2013;15:R102. [Europe PMC free article] [Abstract] [Google Scholar]
- Bae Y, Yang T, Zeng HC. miRNA-34c regulates Notch signaling during bone development. Hum Mol Genet. 2012;21:2991–3000. [Europe PMC free article] [Abstract] [Google Scholar]
- Mizuno Y, Tokuzawa Y, Ninomiya Y. miR-210 promotes osteoblastic differentiation through inhibition of AcvR1b. FEBS Lett. 2009;583:2263–2268. [Abstract] [Google Scholar]
- Mei Y, Bian C, Li J. miR-21 modulates the ERK-MAPK signaling pathway by regulating SPRY2 expression during human mesenchymal stem cell differentiation. J Cell Biochem. 2013;114:1374–1384. [Abstract] [Google Scholar]
- Goff LA, Boucher S, Ricupero CL. Differentiating human multipotent mesenchymal stromal cells regulate microRNAs: prediction of microRNA regulation by PDGF during osteogenesis. Exp Hematol. 2008;36:1354–1369. [Europe PMC free article] [Abstract] [Google Scholar]
- Kim YJ, Bae SW, Yu SS, Bae YC, Jung JS. miR-196a regulates proliferation and osteogenic differentiation in mesenchymal stem cells derived from human adipose tissue. J Bone Miner Res. 2009;24:816–825. [Abstract] [Google Scholar]
- Schoolmeesters A, Eklund T, Leake D. Functional profiling reveals critical role for miRNA in differentiation of human mesenchymal stem cells. PLoS ONE. 2009;4:e5605. [Europe PMC free article] [Abstract] [Google Scholar]
- Kahai S, Lee SC, Lee DY. MicroRNA miR-378 regulates nephronectin expression modulating osteoblast differentiation by targeting GalNT-7. PLoS ONE. 2009;4:e7535. [Europe PMC free article] [Abstract] [Google Scholar]
- Oskowitz A, Lu J, Penfornis P. Human multipotent stromal cells from bone marrow and microRNA: regulation of differentiation and leukemia inhibitory factor expression. Proc Natl Acad Sci U S A. 2008;105:18372–18377. [Europe PMC free article] [Abstract] [Google Scholar]
- Kuo PL, Liao SH, Hung JY, Huang MS, Hsu YL. MicroRNA-33a functions as a bone metastasis suppressor in lung cancer by targeting parathyroid hormone related protein. Biochim Biophys Acta. 2013;1830:3756–3766. [Abstract] [Google Scholar]
- Pollari S, Leivonen SK, Perälä M, Fey V, Käkönen SM, Kallioniemi O. Identification of microRNAs inhibiting TGF-β-induced IL-11 production in bone metastatic breast cancer cells. PLoS ONE. 2012;7:e37361. [Europe PMC free article] [Abstract] [Google Scholar]
- Zhao FL, Hu GD, Wang XF, Zhang XH, Zhang YK, Yu ZS. Serum overexpression of microRNA-10b in patients with bone metastatic primary breast cancer. J Int Med Res. 2012;40:859–866. [Abstract] [Google Scholar]
- Tie J, Pan Y, Zhao L. MiR-218 inhibits invasion and metastasis of gastric cancer by targeting the Robo1 receptor. PLoS Genet. 2010;6:e1000879. [Europe PMC free article] [Abstract] [Google Scholar]
- Octacílio-Silva S, Marques MM, Evangelista AF. Defining mRNA Targets of MicroRNAs during Osteoblastic Differentiation of Human Mesenchymal Stem Cells by Microarray Transcriptional Interaction Networks. Resumos do 56° Congresso Brasileiro de Genética, Guarujá-SP, Brasil; 2010. [Google Scholar]
- Valencia K, Martín-Fernαndez M, Zandueta C. miR-326 associates with biochemical markers of bone turnover in lung cancer bone metastasis. Bone. 2013;52:532–539. [Abstract] [Google Scholar]
- Hassan MQ, Gordon JA, Beloti MM. A network connecting Runx2, SATB2, and the miR-23a~27a~24–2 cluster regulates the osteoblast differentiation program. Proc Natl Acad Sci U S A. 2010;107:19879–19884. [Europe PMC free article] [Abstract] [Google Scholar]
- Zeng Y, Qu X, Li H. MicroRNA-100 regulates osteogenic differentiation of human adipose-derived mesenchymal stem cells by targeting BMPR2. FEBS Lett. 2012;586:2375–2381. [Abstract] [Google Scholar]
- Vimalraj S, Selvamurugan N. MicroRNAs: Synthesis, gene regulation and osteoblast differentiation. Curr Issues Mol Biol. 2012;15:7–18. [Abstract] [Google Scholar]
- Inosea H, Ochi H, Kimura A. A microRNA regulatory mechanism of osteoblast differentiation. Proc Natl Acad Sci U S A. 2009;106:20794–20799. [Europe PMC free article] [Abstract] [Google Scholar]
- Huang G, Nishimoto K, Zhou Z, Hughes D, Kleinerman ES. miR-20a encoded by the miR-17-92 cluster increases the metastatic potential of osteosarcoma cells by regulating Fas expression. Cancer Res. 2012;72:908–916. [Europe PMC free article] [Abstract] [Google Scholar]
- Tomé M, López-Romero P, Albo C. miR-335 orchestrates cell proliferation, migration and differentiation in human mesenchymal stem cells. Cell Death Diff. 2011;18:985–995. [Europe PMC free article] [Abstract] [Google Scholar]
- Yin Q, Wang X, Fewell C. MicroRNA miR-155 inhibits bone morphogenetic protein (BMP) signaling and BMP-mediated Epstein–Barr virus reactivation. J Virol. 2010;84:6318–6327. [Europe PMC free article] [Abstract] [Google Scholar]
- Mizuno Y, Yagi K, Tokuzawa Y. miR-125b inhibits osteoblastic differentiation by down-regulation of cell proliferation. Biochem Biophys Res Commun. 2008;368:267–272. [Abstract] [Google Scholar]
- Chang L, Karin M. Mammalian MAP kinase signalling cascades. Nature. 2001;410:37–40. [Abstract] [Google Scholar]
- Kretzschmar M, Doody J, Timokhina I, Massague J. A mechanism of repression of TGF beta/Smad signaling by oncogenic Ras. Genes Dev. 1999;13:804–816. [Europe PMC free article] [Abstract] [Google Scholar]
- Sapkota G, Alarcon C, Spagnoli FM, Brivanlou AH, Massague J. Balancing BMP signaling through integrated inputs into the Smad1 linker. Mol Cell. 2007;25:441–454. [Abstract] [Google Scholar]
- Matsuura I, Wang G, He D, Liu F. Identification and characterization of ERK MAP kinase phosphorylation sites in Smad3. Biochemistry. 2005;44:12546–12553. [Abstract] [Google Scholar]
- Herpin A, Cunningham C. Cross-talk between the bone morphogenetic protein pathway and other major signaling pathways results in tightly regulated cell-specific outcomes. FEBS J. 2007;274:2977–2985. [Abstract] [Google Scholar]
- Leung-Hagesteijn C, Hu MC, Mahendra AS. Integrin-linked kinase mediates bone morphogenetic protein 7-dependent renal epithelial cell morphogenesis. Mol Cell Biol. 2005;25:3648–3657. [Europe PMC free article] [Abstract] [Google Scholar]
- Eivers E, Fuentealba LC, De Robertis EM. Integrating positional information at the level of Smad1/5/8. Curr Opin Genet Dev. 2008;18:304–310. [Europe PMC free article] [Abstract] [Google Scholar]
- Ghosh-Choudhury N, Abboud SL, Nishimura R, Celeste A, Mahimainathan L, Choudhury GG. Requirement of BMP-2-induced phosphatidylinositol 3-kinase and Akt serine/threonine kinase in osteoblast differentiation and Smad-dependent BMP-2 gene transcription. J Biol Chem. 2002;277:33361–33368. [Abstract] [Google Scholar]
- Moustakas A, Heldin CH. Non-Smad TGF-beta signals. J Cell Sci. 2005;118:3573–3584. [Abstract] [Google Scholar]
- Engin F, Lee B. NOTCHing the bone: insights into multi-functionality. Bone. 2010;46:274–280. [Europe PMC free article] [Abstract] [Google Scholar]
- Tezuka K, Yasuda M, Watanabe N. Stimulation of osteoblastic cell differentiation by Notch. J Bone Miner Res. 2002;17:231–239. [Abstract] [Google Scholar]
- Turnpenny PD, Alman B, Cornier AS. Abnormal vertebral segmentation and the notch signaling pathway in man. Dev Dyn. 2007;236:1456–1474. [Abstract] [Google Scholar]
- Ramasamy SK, Kusumbe AP, Wang L, Adams RH. Endothelial Notch activity promotes angiogenesis and osteogenesis in bone. Nature. 2014;507:376–380. [Europe PMC free article] [Abstract] [Google Scholar]
- Luppen CA, Chandler RL, Noh T, Mortlock DP, Frenkel B. BMP-2 vs. BMP-4 expression and activity in glucocorticoid arrested MC3T3-E1 osteoblasts: Smad signaling, not alkaline phosphatase activity, predicts rescue of mineralization. Growth Factors. 2008;26:226–237. [Europe PMC free article] [Abstract] [Google Scholar]
- Sheng N, Xie Z, Wang C. Retinoic acid regulates bone morphogenic protein signal duration by promoting the degradation of phosphorylated Smad1. Proc Natl Acad Sci U S A. 2010;107:18886–18891. [Europe PMC free article] [Abstract] [Google Scholar]
- McKeown SJ, Wallace AS, Anderson RB. Expression and function of cell adhesion molecules during neural crest migration. Dev Biol. 2013;373:244–257. [Abstract] [Google Scholar]
- Zhou J, Lee PL, Lee CI. BMP receptor-integrin interaction mediates responses of vascular endothelial Smad1/5 and proliferation to disturbed flow. J Thromb Haemost. 2013;11:741–755. [Abstract] [Google Scholar]
- Lamouille S, Connolly E, Smyth JW, Akhurst RJ, Derynck R. TGF-b-induced activation of mTOR complex 2 drives epithelial–mesenchymal transition and cell invasion. J Cell Sci. 2012;125:1259–1273. [Europe PMC free article] [Abstract] [Google Scholar]
- Xu J, Lamouille S, Derynck R. TGF-b induced epithelial to mesenchymal transition. Cell Res. 2009;19:156–172. [Europe PMC free article] [Abstract] [Google Scholar]
- Song K, Wang H, Krebs TL, Danielpour D. Novel roles of Akt and mTOR in suppressing TGF-b/ALK5-mediated Smad3 activation. EMBO J. 2006;25:58–69. [Europe PMC free article] [Abstract] [Google Scholar]
- Dreieicher E, Beck KF, Lazaroski S. Nitric oxide inhibits glomerular TGF-b signaling via SMOC-1. J Am Soc Nephrol. 2009;20:1963–1974. [Europe PMC free article] [Abstract] [Google Scholar]
- Saura M, Zaragoza C, Herranz B. Nitric oxide regulates transforming growth factor-β signaling in endothelial Cells. Circ Res. 2005;97:1115–1123. [Abstract] [Google Scholar]
- Saito T, Fukai A, Mabuchi A. Transcriptional regulation of endochondral ossification by HIF-2alpha during skeletal growth and osteoarthritis development. Nat Med. 2010;16:678–686. [Abstract] [Google Scholar]
- Mariani E, Pulsatelli L, Facchini A. Signaling pathways in cartilage repair. Int J Mol Sci. 2014;15:8667–8698. [Europe PMC free article] [Abstract] [Google Scholar]
- Sagar DR, Ashraf S, Xu L. Osteoprotegerin reduces the development of pain behaviour and joint pathology in a model of osteoarthritis. Ann Rheum Dis. 2014;73:1558–1565. [Europe PMC free article] [Abstract] [Google Scholar]
- Nakashima T, Takayanagi H. The dynamic interplay between osteoclasts and the immune system. Arch Biochem Biophys. 2008;473:166–171. [Abstract] [Google Scholar]
- Jimi E, Hirata S, Shin M, Yamazaki M, Fukushima H. Molecular mechanisms of BMP-induced bone formation: cross-talk between BMP and NF-κB signaling pathways in osteoblastogenesis. Jpn Dent Sci Rev. 2010;46:33–42. [Google Scholar]
- Eliseev RA, Schwarz EM, Zuscik MJ, O'Keefe RJ, Drissi H, Rosier RN. Smad7 mediates inhibition of Saos2 osteosarcoma cell differentiation by NF-κB. Exp Cell Res. 2006;312:40–50. [Abstract] [Google Scholar]
- Li Y, Li A, Strait K, Zhang H, Nanes MS, Weitzmann MN. Endogenous TNFa lowers maximum peak bone mass and inhibits osteoblastic Smad activation through NF-κB. J Bone Miner Res. 2007;22:646–655. [Abstract] [Google Scholar]
- Kramer I, Halleux C, Feng JQ, Bonewald LF, Kneissel M. Osteocyte Wnt/β-catenin signaling is required for normal bone homeostasis. Mol Cell Biol. 2010;30:3071–3085. [Europe PMC free article] [Abstract] [Google Scholar]
- Zhang Y, Khan D, Delling J, Tobiasch E. Mechanisms underlying the osteo and adipo differentiation of human mesenchymal stem cells. Sci World J. 2012;2012:793823. [Europe PMC free article] [Abstract] [Google Scholar]
- Das S, Samant RS, Shevde LA. The hedgehog pathway conditions the bone microenvironment for osteolytic metastasis of breast cancer. Int J Breast Cancer. 2012;2012:298623. [Europe PMC free article] [Abstract] [Google Scholar]
- Hughes FJ, Turner W, G Belibasakis G, Martuscelli G. Effects of growth factors and cytokines on osteoblast differentiation. Periodontol 2000. 2006;41:48–72. [Abstract] [Google Scholar]
- Fuller K, Lawrence KM, Ross JL. Cathepsin K inhibitors prevent matrix-derived growth factor degradation by human osteoclasts. Bone. 2008;42:200–211. [Abstract] [Google Scholar]
- Xie H, Cui Z, Wang L. PDGF-BB secreted by preosteoclasts induces angiogenesis during coupling with osteogenesis. Nat Med. 2014;20:1270–1278. [Europe PMC free article] [Abstract] [Google Scholar]
- Gerstenfeld L, Edgar CM, Kakar S, Jacobsen KA, Einhorn TA. Osteogenic growth factors and cytokines and their role in bone repair engineering of functional skeletal tissues. Topics Bone Biol. 2007;3:17–45. [Google Scholar]
- Li X, Cao X. BMP signaling and skeletogenesis. Ann N Y Acad Sci. 2006;1068:26–40. [Abstract] [Google Scholar]
- Shimizu T, Tanaka T, Iso T. Notch signaling pathway enhances bone morphogenetic protein 2 (BMP2) responsiveness of Msx2 gene to induce osteogenic differentiation and mineralization of vascular smooth muscle cells. J Biol Chem. 2011;286:19138–19148. [Europe PMC free article] [Abstract] [Google Scholar]
- Sowa H, Kaji H, Hendy GN. Menin is required for bone morphogenetic protein 2- and transforming growth factor beta-regulated osteoblastic differentiation through interaction with Smads and Runx2. J Biol Chem. 2004;279:40267–40275. [Abstract] [Google Scholar]
- Li X, Cao X. BMP signaling and HOX transcription factors in limb development. Front Biosci. 2003;8:s805–s812. [Abstract] [Google Scholar]
Articles from Bone Research are provided here courtesy of Nature Publishing Group
Full text links
Read article at publisher's site: https://doi.org/10.1038/boneres.2015.5
Read article for free, from open access legal sources, via Unpaywall:
https://www.nature.com/articles/boneres20155.pdf
Citations & impact
Impact metrics
Citations of article over time
Alternative metrics
Smart citations by scite.ai
Explore citation contexts and check if this article has been
supported or disputed.
https://scite.ai/reports/10.1038/boneres.2015.5
Article citations
Molecular Signaling Pathways and MicroRNAs in Bone Remodeling: A Narrative Review.
Diseases, 12(10):252, 12 Oct 2024
Cited by: 0 articles | PMID: 39452495 | PMCID: PMC11507001
Review Free full text in Europe PMC
Multifunctional vitamin D-incorporated PLGA scaffold with BMP/VEGF-overexpressed tonsil-derived MSC via CRISPR/Cas9 for bone tissue regeneration.
Mater Today Bio, 28:101254, 14 Sep 2024
Cited by: 0 articles | PMID: 39328787 | PMCID: PMC11426062
Emerging role of BMPs/BMPR2 signaling pathway in treatment for pulmonary fibrosis.
Biomed Pharmacother, 178:117178, 13 Aug 2024
Cited by: 0 articles | PMID: 39142248 | PMCID: PMC11364484
Review Free full text in Europe PMC
The Role of Circular RNA in the Pathogenesis of Chemotherapy-Induced Cardiotoxicity in Cancer Patients: Focus on the Pathogenesis and Future Perspective.
Cardiovasc Toxicol, 24(11):1151-1167, 19 Aug 2024
Cited by: 0 articles | PMID: 39158829
Review
Apoptosis and Inflammation Involved with Fluoride-Induced Bone Injuries.
Nutrients, 16(15):2500, 31 Jul 2024
Cited by: 0 articles | PMID: 39125380 | PMCID: PMC11313706
Go to all (292) article citations
Data
Data behind the article
This data has been text mined from the article, or deposited into data resources.
BioStudies: supplemental material and supporting data
Genes & Proteins
- (1 citation) UniProt - P18075
Protein structures in PDBe (3)
-
(2 citations)
PDBe - 2GOOView structure
-
(1 citation)
PDBe - 3BMPView structure
-
(1 citation)
PDBe - 1MHDView structure
Similar Articles
To arrive at the top five similar articles we use a word-weighted algorithm to compare words from the Title and Abstract of each citation.
TGF-β and BMP signaling in osteoblast, skeletal development, and bone formation, homeostasis and disease.
Bone Res, 4:16009, 26 Apr 2016
Cited by: 739 articles | PMID: 27563484 | PMCID: PMC4985055
Review Free full text in Europe PMC
Bone morphogenetic proteins.
Growth Factors, 22(4):233-241, 01 Dec 2004
Cited by: 1246 articles | PMID: 15621726
Review
Localization of Smads, the TGF-beta family intracellular signaling components during endochondral ossification.
J Bone Miner Res, 14(7):1145-1152, 01 Jul 1999
Cited by: 95 articles | PMID: 10404014
Divergence and convergence of TGF-beta/BMP signaling.
J Cell Physiol, 187(3):265-276, 01 Jun 2001
Cited by: 314 articles | PMID: 11319750
Review